+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7869 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mu Opioid Receptor-Gi Protein Complex | ||||||||||||||||||
 Map data Map data | Mu Opioid Receptor-Gi Protein Complex, full map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Complex / Transmembrane / MEMBRANE PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationOpioid Signalling / spine apparatus / positive regulation of appetite / sperm ejaculation / G-protein activation / Peptide ligand-binding receptors / beta-endorphin receptor activity / morphine receptor activity / negative regulation of Wnt protein secretion / adenylate cyclase-inhibiting opioid receptor signaling pathway ...Opioid Signalling / spine apparatus / positive regulation of appetite / sperm ejaculation / G-protein activation / Peptide ligand-binding receptors / beta-endorphin receptor activity / morphine receptor activity / negative regulation of Wnt protein secretion / adenylate cyclase-inhibiting opioid receptor signaling pathway / negative regulation of luteinizing hormone secretion / G protein-coupled opioid receptor activity / regulation of cellular response to stress / G protein-coupled opioid receptor signaling pathway / G alpha (i) signalling events / negative regulation of nitric oxide biosynthetic process / negative regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / behavioral response to ethanol / regulation of NMDA receptor activity / neuropeptide binding / positive regulation of neurogenesis / eating behavior / negative regulation of cytosolic calcium ion concentration / transmission of nerve impulse / G-protein alpha-subunit binding / social behavior / voltage-gated calcium channel activity / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / T-tubule / sensory perception of pain / positive regulation of gluconeogenesis / presynaptic modulation of chemical synaptic transmission / dendrite membrane / dendrite cytoplasm / cellular response to forskolin / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / sarcoplasmic reticulum / Regulation of insulin secretion / neuropeptide signaling pathway / locomotory behavior / excitatory postsynaptic potential / response to prostaglandin E / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / sarcolemma / G protein-coupled receptor binding / response to peptide hormone / G protein-coupled receptor activity / GABA-ergic synapse / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / GDP binding / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G alpha (12/13) signalling events / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||||||||
 Authors Authors | Koehl A / Hu H | ||||||||||||||||||
| Funding support |  United States, United States,  Switzerland, 5 items Switzerland, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Structure of the µ-opioid receptor-G protein complex. Authors: Antoine Koehl / Hongli Hu / Shoji Maeda / Yan Zhang / Qianhui Qu / Joseph M Paggi / Naomi R Latorraca / Daniel Hilger / Roger Dawson / Hugues Matile / Gebhard F X Schertler / Sebastien ...Authors: Antoine Koehl / Hongli Hu / Shoji Maeda / Yan Zhang / Qianhui Qu / Joseph M Paggi / Naomi R Latorraca / Daniel Hilger / Roger Dawson / Hugues Matile / Gebhard F X Schertler / Sebastien Granier / William I Weis / Ron O Dror / Aashish Manglik / Georgios Skiniotis / Brian K Kobilka /    Abstract: The μ-opioid receptor (μOR) is a G-protein-coupled receptor (GPCR) and the target of most clinically and recreationally used opioids. The induced positive effects of analgesia and euphoria are ...The μ-opioid receptor (μOR) is a G-protein-coupled receptor (GPCR) and the target of most clinically and recreationally used opioids. The induced positive effects of analgesia and euphoria are mediated by μOR signalling through the adenylyl cyclase-inhibiting heterotrimeric G protein G. Here we present the 3.5 Å resolution cryo-electron microscopy structure of the μOR bound to the agonist peptide DAMGO and nucleotide-free G. DAMGO occupies the morphinan ligand pocket, with its N terminus interacting with conserved receptor residues and its C terminus engaging regions important for opioid-ligand selectivity. Comparison of the μOR-G complex to previously determined structures of other GPCRs bound to the stimulatory G protein G reveals differences in the position of transmembrane receptor helix 6 and in the interactions between the G protein α-subunit and the receptor core. Together, these results shed light on the structural features that contribute to the G protein-coupling specificity of the µOR. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7869.map.gz emd_7869.map.gz | 48.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7869-v30.xml emd-7869-v30.xml emd-7869.xml emd-7869.xml | 24.6 KB 24.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7869.png emd_7869.png | 58.7 KB | ||
| Filedesc metadata |  emd-7869.cif.gz emd-7869.cif.gz | 7.3 KB | ||
| Others |  emd_7869_half_map_1.map.gz emd_7869_half_map_1.map.gz emd_7869_half_map_2.map.gz emd_7869_half_map_2.map.gz | 49.4 MB 49.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7869 http://ftp.pdbj.org/pub/emdb/structures/EMD-7869 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7869 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7869 | HTTPS FTP |
-Related structure data
| Related structure data |  6ddfMC  7868C  6ddeC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7869.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7869.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mu Opioid Receptor-Gi Protein Complex, full map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
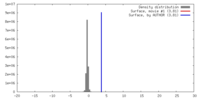
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Mu Opioid Receptor-Gi Protein Complex, half map #1
| File | emd_7869_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mu Opioid Receptor-Gi Protein Complex, half map #1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
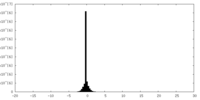
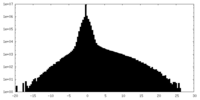
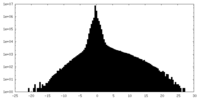
| Density Histograms |
-Half map: Mu Opioid Receptor-Gi Protein Complex, half map #2
| File | emd_7869_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mu Opioid Receptor-Gi Protein Complex, half map #2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
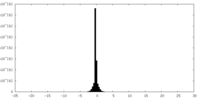
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of DAMGO-activated Mu-type opioid receptor with h...
| Entire | Name: Ternary complex of DAMGO-activated Mu-type opioid receptor with heterotrimeric Gi |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of DAMGO-activated Mu-type opioid receptor with h...
| Supramolecule | Name: Ternary complex of DAMGO-activated Mu-type opioid receptor with heterotrimeric Gi type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Signaling complex formed by incubation of DAMGO-bound Mu-type opioid receptor and heterotrimeric Gi. Excess GDP removed by addition of apyrase. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 120 KDa |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.415031 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGGQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCA TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.671102 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: PGSSGSELDQ LRQEAEQLKN QIRDARKACA DATLSQITNN IDPVGRIQMR TRRTLRGHLA KIYAMHWGTD SRLLVSASQD GKLIIWDSY TTNKVHAIPL RSSWVMTCAY APSGNYVACG GLDNICSIYN LKTREGNVRV SRELAGHTGY LSCCRFLDDN Q IVTSSGDT ...String: PGSSGSELDQ LRQEAEQLKN QIRDARKACA DATLSQITNN IDPVGRIQMR TRRTLRGHLA KIYAMHWGTD SRLLVSASQD GKLIIWDSY TTNKVHAIPL RSSWVMTCAY APSGNYVACG GLDNICSIYN LKTREGNVRV SRELAGHTGY LSCCRFLDDN Q IVTSSGDT TCALWDIETG QQTTTFTGHT GDVMSLSLAP DTRLFVSGAC DASAKLWDVR EGMCRQTFTG HESDINAICF FP NGNAFAT GSDDATCRLF DLRADQELMT YSHDNIICGI TSVSFSKSGR LLLAGYDDFN CNVWDALKAD RAGVLAGHDN RVS CLGVTD DGMAVATGSW DSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Mu-type opioid receptor
| Macromolecule | Name: Mu-type opioid receptor / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.995105 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: NISDCSDPLA PASCSPAPGS WLNLSHVDGN QSDPCGPNRT GLGENLYFQG SHSLCPQTGS PSMVTAITIM ALYSIVCVVG LFGNFLVMY VIVRYTKMKT ATNIYIFNLA LADALATSTL PFQSVNYLMG TWPFGNILCK IVISIDYYNM FTSIFTLCTM S VDRYIAVC ...String: NISDCSDPLA PASCSPAPGS WLNLSHVDGN QSDPCGPNRT GLGENLYFQG SHSLCPQTGS PSMVTAITIM ALYSIVCVVG LFGNFLVMY VIVRYTKMKT ATNIYIFNLA LADALATSTL PFQSVNYLMG TWPFGNILCK IVISIDYYNM FTSIFTLCTM S VDRYIAVC HPVKALDFRT PRNAKIVNVC NWILSSAIGL PVMFMATTKY RQGSIDCTLT FSHPTWYWEN LLKICVFIFA FI MPVLIIT VCYGLMILRL KSVRMLSGSK EKDRNLRRIT RMVLVVVAVF IVCWTPIHIY VIIKALITIP ETTFQTVSWH FCI ALGYTN SCLNPVLYAF LDENFKRCFR EFCIPTSSTI UniProtKB: Mu-type opioid receptor |
-Macromolecule #5: DAMGO
| Macromolecule | Name: DAMGO / type: protein_or_peptide / ID: 5 / Details: analogue of enkephalin / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 513.587 Da |
| Sequence | String: Y(DAL)G(MEA)(ETA) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7 mg/mL | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: Solutions were made fresh. After buffer reconstitution, all buffers were filtered with 0.22um filter. | ||||||||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa | ||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV / Details: blot 1 second before plunging. | ||||||||||||||||||||||||
| Details | This sample was monodisperse by negative stain and Cryo-EM analysis. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 2642 / Average exposure time: 8.0 sec. / Average electron dose: 37.0 e/Å2 Details: Images were collected in movie-mode at 10 frames per second. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 48076 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)