+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7123 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of rat TRPV6* in nanodiscs | |||||||||
 Map data Map data | primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion channel / Membrane protein / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationparathyroid hormone secretion / TRP channels / calcium ion import / calcium-activated cation channel activity / calcium ion import across plasma membrane / calcium ion homeostasis / calcium channel complex / response to calcium ion / calcium ion transmembrane transport / calcium channel activity ...parathyroid hormone secretion / TRP channels / calcium ion import / calcium-activated cation channel activity / calcium ion import across plasma membrane / calcium ion homeostasis / calcium channel complex / response to calcium ion / calcium ion transmembrane transport / calcium channel activity / calcium ion transport / protein homotetramerization / calmodulin binding / apical plasma membrane / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | McGoldrick LL / Singh AK / Saotome K / Yelshanskaya MV / Twomey EC / Grassucci RA / Sobolevsky AI | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Opening of the human epithelial calcium channel TRPV6. Authors: Luke L McGoldrick / Appu K Singh / Kei Saotome / Maria V Yelshanskaya / Edward C Twomey / Robert A Grassucci / Alexander I Sobolevsky /  Abstract: Calcium-selective transient receptor potential vanilloid subfamily member 6 (TRPV6) channels play a critical role in calcium uptake in epithelial tissues. Altered TRPV6 expression is associated with ...Calcium-selective transient receptor potential vanilloid subfamily member 6 (TRPV6) channels play a critical role in calcium uptake in epithelial tissues. Altered TRPV6 expression is associated with a variety of human diseases, including cancers. TRPV6 channels are constitutively active and their open probability depends on the lipidic composition of the membrane in which they reside; it increases substantially in the presence of phosphatidylinositol 4,5-bisphosphate. Crystal structures of detergent-solubilized rat TRPV6 in the closed state have previously been solved. Corroborating electrophysiological results, these structures demonstrated that the Ca selectivity of TRPV6 arises from a ring of aspartate side chains in the selectivity filter that binds Ca tightly. However, how TRPV6 channels open and close their pores for ion permeation has remained unclear. Here we present cryo-electron microscopy structures of human TRPV6 in the open and closed states. The channel selectivity filter adopts similar conformations in both states, consistent with its explicit role in ion permeation. The iris-like channel opening is accompanied by an α-to-π-helical transition in the pore-lining transmembrane helix S6 at an alanine hinge just below the selectivity filter. As a result of this transition, the S6 helices bend and rotate, exposing different residues to the ion channel pore in the open and closed states. This gating mechanism, which defines the constitutive activity of TRPV6, is, to our knowledge, unique among tetrameric ion channels and provides structural insights for understanding their diverse roles in physiology and disease. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7123.map.gz emd_7123.map.gz | 5.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7123-v30.xml emd-7123-v30.xml emd-7123.xml emd-7123.xml | 17.6 KB 17.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7123.png emd_7123.png | 96 KB | ||
| Filedesc metadata |  emd-7123.cif.gz emd-7123.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7123 http://ftp.pdbj.org/pub/emdb/structures/EMD-7123 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7123 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7123 | HTTPS FTP |
-Related structure data
| Related structure data |  6bobMC  7120C  7121C  7122C  6bo8C  6bo9C  6boaC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7123.map.gz / Format: CCP4 / Size: 32.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7123.map.gz / Format: CCP4 / Size: 32.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.98 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
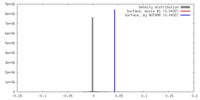
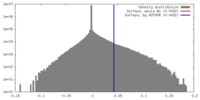
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : TRPV6* ion channel in closed states
| Entire | Name: TRPV6* ion channel in closed states |
|---|---|
| Components |
|
-Supramolecule #1: TRPV6* ion channel in closed states
| Supramolecule | Name: TRPV6* ion channel in closed states / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 6
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 6 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 77.133492 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSLPKEKG LILCLWNKFC RWFHRRESWA QSRDEQNLLQ QKRIWESPLL LAAKENNVQA LYKLLKFEGC EVHQKGAMGE TALHIAALY DNNEAAQVLM EAAPELVFEP MTSELYEGQT ALHIAVINQN VNLVRALLAR GASVSARATG SVFHYRPHNL I YYGEHPLS ...String: MGWSLPKEKG LILCLWNKFC RWFHRRESWA QSRDEQNLLQ QKRIWESPLL LAAKENNVQA LYKLLKFEGC EVHQKGAMGE TALHIAALY DNNEAAQVLM EAAPELVFEP MTSELYEGQT ALHIAVINQN VNLVRALLAR GASVSARATG SVFHYRPHNL I YYGEHPLS FAACVGSEEI VRLLIEHGAD IRAQDSLGNT VLHILILQPN KTFACQMYNL LLSYDGGDHL KSLELVPNNQ GL TPFKLAG VEGNIVMFQH LMQKRKHIQW TYGPLTSTLY DLTEIDSSGD DQSLLELIVT TKKREARQIL DQTPVKELVS LKW KRYGRP YFCVLGAIYV LYIICFTMCC VYRPLKPRIT NRTNPRDNTL LQQKLLQEAY VTPKDDLRLV GELVSIVGAV IILL VEIPD IFRLGVTRFF GQTILGGPFH VIIVTYAFMV LVTMVMRLTN SDGEVVPMSF ALVLGWCNVM YFARGFQMLG PFTIM IQKM IFGDLMRFCW LMAVVILGFA SAFYIIFQTE DPDELGHFYD YPMALFSTFE LFLTIIDGPA NYDVDLPFMY SITYAA FAI IATLLMLNLL IAMMGDTHWR VAHERDELWR AQVVATTVML ERKLPRCLWP RSGICGREYG LGDRWFLRVE DRQDLNR QR IRRYAQAFQQ QDDLYSEDLE KDSGEKLVPR UniProtKB: Transient receptor potential cation channel subfamily V member 6 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Component: (Formula: TRIS, NaCl) / Details: TRIS, NaCl, |
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6bob: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)