[English] 日本語
 Yorodumi
Yorodumi- EMDB-7082: cryo-EM structure of TRPM4 in apo state with long coiled coil at ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7082 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of TRPM4 in apo state with long coiled coil at 3.5 angstrom resolution | ||||||||||||||||||
 Map data Map data | Structure of TRPM4 in apo state with long coiled coil at 3.5 angstrom | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | ion channel / TRANSPORT PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of atrial cardiac muscle cell action potential / positive regulation of regulation of vascular associated smooth muscle cell membrane depolarization / sodium channel complex / regulation of T cell cytokine production / membrane depolarization during AV node cell action potential / membrane depolarization during bundle of His cell action potential / membrane depolarization during Purkinje myocyte cell action potential / negative regulation of bone mineralization / metal ion transport / TRP channels ...positive regulation of atrial cardiac muscle cell action potential / positive regulation of regulation of vascular associated smooth muscle cell membrane depolarization / sodium channel complex / regulation of T cell cytokine production / membrane depolarization during AV node cell action potential / membrane depolarization during bundle of His cell action potential / membrane depolarization during Purkinje myocyte cell action potential / negative regulation of bone mineralization / metal ion transport / TRP channels / voltage-gated monoatomic ion channel activity / regulation of ventricular cardiac muscle cell action potential / calcium-activated cation channel activity / sodium ion import across plasma membrane / : / dendritic cell chemotaxis / cellular response to ATP / regulation of heart rate by cardiac conduction / monoatomic cation transmembrane transport / protein sumoylation / positive regulation of vasoconstriction / negative regulation of osteoblast differentiation / positive regulation of fat cell differentiation / positive regulation of heart rate / long-term memory / positive regulation of adipose tissue development / positive regulation of insulin secretion involved in cellular response to glucose stimulus / calcium-mediated signaling / regulation of membrane potential / calcium channel activity / calcium ion transport / positive regulation of canonical Wnt signaling pathway / positive regulation of cytosolic calcium ion concentration / protein homotetramerization / adaptive immune response / calmodulin binding / neuronal cell body / calcium ion binding / positive regulation of cell population proliferation / endoplasmic reticulum / Golgi apparatus / nucleoplasm / ATP binding / membrane / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.54 Å | ||||||||||||||||||
 Authors Authors | Guo J / She J | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Structures of the calcium-activated, non-selective cation channel TRPM4. Authors: Jiangtao Guo / Ji She / Weizhong Zeng / Qingfeng Chen / Xiao-Chen Bai / Youxing Jiang /  Abstract: TRPM4 is a calcium-activated, phosphatidylinositol-4,5-bisphosphate (PtdIns(4,5)P) -modulated, non-selective cation channel that belongs to the family of melastatin-related transient receptor ...TRPM4 is a calcium-activated, phosphatidylinositol-4,5-bisphosphate (PtdIns(4,5)P) -modulated, non-selective cation channel that belongs to the family of melastatin-related transient receptor potential (TRPM) channels. Here we present the electron cryo-microscopy structures of the mouse TRPM4 channel with and without ATP. TRPM4 consists of multiple transmembrane and cytosolic domains, which assemble into a three-tiered architecture. The N-terminal nucleotide-binding domain and the C-terminal coiled-coil participate in the tetrameric assembly of the channel; ATP binds at the nucleotide-binding domain and inhibits channel activity. TRPM4 has an exceptionally wide filter but is only permeable to monovalent cations; filter residue Gln973 is essential in defining monovalent selectivity. The S1-S4 domain and the post-S6 TRP domain form the central gating apparatus that probably houses the Ca- and PtdIns(4,5)P-binding sites. These structures provide an essential starting point for elucidating the complex gating mechanisms of TRPM4 and reveal the molecular architecture of the TRPM family. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7082.map.gz emd_7082.map.gz | 60.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7082-v30.xml emd-7082-v30.xml emd-7082.xml emd-7082.xml | 22.5 KB 22.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7082.png emd_7082.png | 207.1 KB | ||
| Filedesc metadata |  emd-7082.cif.gz emd-7082.cif.gz | 7.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7082 http://ftp.pdbj.org/pub/emdb/structures/EMD-7082 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7082 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7082 | HTTPS FTP |
-Related structure data
| Related structure data |  6bclMC  7081C  7083C  7085C  6bcjC  6bcoC  6bcqC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7082.map.gz / Format: CCP4 / Size: 65.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7082.map.gz / Format: CCP4 / Size: 65.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of TRPM4 in apo state with long coiled coil at 3.5 angstrom | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
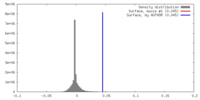
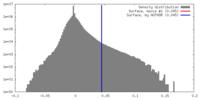
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : homotetramer of mouse TRPM4
| Entire | Name: homotetramer of mouse TRPM4 |
|---|---|
| Components |
|
-Supramolecule #1: homotetramer of mouse TRPM4
| Supramolecule | Name: homotetramer of mouse TRPM4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 4
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 4 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 140.922875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MVGPEKEQSW IPKIFRKKVC TTFIVDLSDD AGGTLCQCGQ PRDAHPSVAV EDAFGAAVVT EWNSDEHTTE KPTDAYGDLD FTYSGRKHS NFLRLSDRTD PATVYSLVTR SWGFRAPNLV VSVLGGSGGP VLQTWLQDLL RRGLVRAAQS TGAWIVTGGL H TGIGRHVG ...String: MVGPEKEQSW IPKIFRKKVC TTFIVDLSDD AGGTLCQCGQ PRDAHPSVAV EDAFGAAVVT EWNSDEHTTE KPTDAYGDLD FTYSGRKHS NFLRLSDRTD PATVYSLVTR SWGFRAPNLV VSVLGGSGGP VLQTWLQDLL RRGLVRAAQS TGAWIVTGGL H TGIGRHVG VAVRDHQTAS TGSSKVVAMG VAPWGVVRNR DMLINPKGSF PARYRWRGDP EDGVEFPLDY NYSAFFLVDD GT YGRLGGE NRFRLRFESY VAQQKTGVGG TGIDIPVLLL LIDGDEKMLK RIEDATQAQL PCLLVAGSGG AADCLVETLE DTL APGSGG LRRGEARDRI RRYFPKGDPE VLQAQVERIM TRKELLTVYS SEDGSEEFET IVLRALVKAC GSSEASAYLD ELRL AVAWN RVDIAQSELF RGDIQWRSFH LEASLMDALL NDRPEFVRLL ISHGLSLGHF LTPVRLAQLY SAVSPNSLIR NLLDQ ASHA SSSKSPPVNG TVELRPPNVG QVLRTLLGET CAPRYPARNT RDSYLGQDHR ENDSLLMDWA NKQPSTDASF EQAPWS DLL IWALLLNRAQ MAIYFWEKGS NSVASALGAC LLLRVMARLE SEAEEAARRK DLAATFESMS VDLFGECYHN SEERAAR LL LRRCPLWGEA TCLQLAMQAD ARAFFAQDGV QSLLTQKWWG EMDSTTPIWA LLLAFFCPPL IYTNLIVFRK SEEEPTQK D LDFDMDSSIN GAGPPGTVEP SAKVALERRQ RRRPGRALCC GKFSKRWSDF WGAPVTAFLG NVVSYLLFLL LFAHVLLVD FQPTKPSVSE LLLYFWAFTL LCEELRQGLG GGWGSLASGG RGPDRAPLRH RLHLYLSDTW NQCDLLALTC FLLGVGCRLT PGLFDLGRT VLCLDFMIFT LRLLHIFTVN KQLGPKIVIV SKMMKDVFFF LFFLCVWLVA YGVATEGILR PQDRSLPSIL R RVFYRPYL QIFGQIPQEE MDVALMIPGN CSMERGSWAH PEGPVAGSCV SQYANWLVVL LLIVFLLVAN ILLLNLLIAM FS YTFSKVH GNSDLYWKAQ RYSLIREFHS RPALAPPLII ISHVRLLIKW LRRCRRCRRA NLPASPVFEH FRVCLSKEAE RKL LTWESV HKENFLLAQA RDKRDSDSER LKRTSQKVDT ALKQLGQIRE YDRRLRGLER EVQHCSRVLT WMAEALSHSA LLPP GAPPP PSPTGSKDRN SKAYVDELTS RGRLEVLFQG PDYKDDDDKH HHHHHHHHH UniProtKB: Transient receptor potential cation channel subfamily M member 4 |
-Macromolecule #2: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 2 / Number of copies: 2 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Component:
| ||||||
|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 80 sec. | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | ||||||
| Details | protein samples are reconstituted into nanodiscs |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Lower energy threshold: -10 eV / Energy filter - Upper energy threshold: 10 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 3000 / Average exposure time: 15.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 46730 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)