+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7070 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
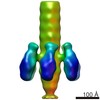
| Title | Structure of the tail of the marine siphovirus TW1 | |||||||||
 Map data Map data | Whole Tail of TW1 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Pseudoalteromonas phage TW1 (virus) Pseudoalteromonas phage TW1 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 23.5 Å | |||||||||
 Authors Authors | Wang Z / Rossmann MG | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2018 Journal: Structure / Year: 2018Title: Structure of the Marine Siphovirus TW1: Evolution of Capsid-Stabilizing Proteins and Tail Spikes. Authors: Zhiqing Wang / Stephen C Hardies / Andrei Fokine / Thomas Klose / Wen Jiang / Byung Cheol Cho / Michael G Rossmann /   Abstract: Marine bacteriophage TW1 belongs to the Siphoviridae family and infects Pseudoalteromonas phenolica. Mass spectrometry analysis has identified 16 different proteins in the TW1 virion. Functions of ...Marine bacteriophage TW1 belongs to the Siphoviridae family and infects Pseudoalteromonas phenolica. Mass spectrometry analysis has identified 16 different proteins in the TW1 virion. Functions of most of these proteins have been predicted by bioinformatic methods. A 3.6 Å resolution cryoelectron microscopy map of the icosahedrally averaged TW1 head showed the atomic structures of the major capsid protein, gp57, and the capsid-stabilizing protein, gp56. The gp57 structure is similar to that of the phage HK97 capsid protein. The gp56 protein has two domains, each having folds similar to that of the N-terminal part of phage λ gpD, indicating a common ancestry. The first gp56 domain clamps adjacent capsomers together, whereas the second domain is required for trimerization. A 6-fold-averaged reconstruction of the distal part of the tail showed that TW1 has six tail spikes, which are unusual for siphophages but are similar to the podophages P22 and Sf6, suggesting a common evolutionary origin of these spikes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7070.map.gz emd_7070.map.gz | 114.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7070-v30.xml emd-7070-v30.xml emd-7070.xml emd-7070.xml | 11.9 KB 11.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7070.png emd_7070.png | 28.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7070 http://ftp.pdbj.org/pub/emdb/structures/EMD-7070 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7070 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7070 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7070.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7070.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Whole Tail of TW1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Pseudoalteromonas phage TW1
| Entire | Name:  Pseudoalteromonas phage TW1 (virus) Pseudoalteromonas phage TW1 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Pseudoalteromonas phage TW1
| Supramolecule | Name: Pseudoalteromonas phage TW1 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / NCBI-ID: 1366055 / Sci species name: Pseudoalteromonas phage TW1 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Virus shell | Shell ID: 1 / Diameter: 620.0 Å / T number (triangulation number): 7 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
Details: 50 mM Tris, pH 7.5, 100 mM NaCl, 8 mM MgSO4 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Mesh: 200 / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 298 K / Instrument: GATAN CRYOPLUNGE 3 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80.0 K / Max: 80.0 K |
| Image recording | Film or detector model: OTHER / Detector mode: COUNTING / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 6.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 37000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















