+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Surface Tubular Element of Vaccinia Virus | |||||||||
 Map data Map data | Surface tubular element(STE) of Vaccinia virus. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | poxvirus / membrane protein / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA-templated transcription termination / helicase activity / hydrolase activity / viral envelope / virion membrane / DNA binding / ATP binding / membrane Similarity search - Function | |||||||||
| Biological species |  Vaccinia virus / Vaccinia virus /  Vaccinia virus (strain Tian Tan) Vaccinia virus (strain Tian Tan) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.23 Å | |||||||||
 Authors Authors | Yu F / Jin G / Liu Y / Sun Z / Lou Z | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Mbio / Year: 2026 Journal: Mbio / Year: 2026Title: Surface Tubular Element of Vaccinia Virus Authors: Yu F / Jin G / Liu Y / Sun Z / Lou Z | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63964.map.gz emd_63964.map.gz | 317.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63964-v30.xml emd-63964-v30.xml emd-63964.xml emd-63964.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_63964.png emd_63964.png | 90.8 KB | ||
| Filedesc metadata |  emd-63964.cif.gz emd-63964.cif.gz | 5.4 KB | ||
| Others |  emd_63964_half_map_1.map.gz emd_63964_half_map_1.map.gz emd_63964_half_map_2.map.gz emd_63964_half_map_2.map.gz | 323.8 MB 317.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63964 http://ftp.pdbj.org/pub/emdb/structures/EMD-63964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63964 | HTTPS FTP |
-Related structure data
| Related structure data |  9u9hMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_63964.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63964.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Surface tubular element(STE) of Vaccinia virus. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map of STE.
| File | emd_63964_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of STE. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map of STE.
| File | emd_63964_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of STE. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Surface Tubular Element of Vaccinia Virus
| Entire | Name: Surface Tubular Element of Vaccinia Virus |
|---|---|
| Components |
|
-Supramolecule #1: Surface Tubular Element of Vaccinia Virus
| Supramolecule | Name: Surface Tubular Element of Vaccinia Virus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Vaccinia virus / Strain: Non-replicating vaccinia virus TianTan Strain Vaccinia virus / Strain: Non-replicating vaccinia virus TianTan Strain |
-Macromolecule #1: Virion membrane protein OPG140
| Macromolecule | Name: Virion membrane protein OPG140 / type: protein_or_peptide / ID: 1 Details: Sequence reference for source organism Vaccinia virus (strain Tian Tan) is not available in UniProt at the time of biocuration. Current sequence reference is from UniProt id P20991. Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Vaccinia virus (strain Tian Tan) Vaccinia virus (strain Tian Tan) |
| Molecular weight | Theoretical: 10.003072 KDa |
| Sequence | String: MDMMLMIGNY FSGVLIAGII LLILSCIFAF IDFSKSTSPT RTWKVLSIMA FILGIIITVG MLIYSMWGKH CAPHRVSGVI HTNHSDISM N UniProtKB: Virion membrane protein OPG140 |
-Macromolecule #2: Mature 21 kDa protein OPG144
| Macromolecule | Name: Mature 21 kDa protein OPG144 / type: protein_or_peptide / ID: 2 Details: Sequence reference for source organism Vaccinia virus (strain Tian Tan) is not available in UniProt at the time of biocuration. Current sequence reference is from UniProt id P68592. Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Vaccinia virus (strain Tian Tan) Vaccinia virus (strain Tian Tan) |
| Molecular weight | Theoretical: 19.027289 KDa |
| Sequence | String: AGVLDKDLFT EEQQQSFMPK DGGMMQNDYG GMNDYLGIFK NNDVRTLLGL ILFVLALYSP PLISILMIFI SSFLLPLTSL VITYCLVTQ MYRGGNGNTV GMSIVCIVAA VIIMAINVFT NSQIFNIISY IILFILFFAY VMNIERQDYR RSINVTIPEQ Y TCNKPYTA G UniProtKB: Virion membrane protein OPG144 precursor |
-Macromolecule #3: 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 3 / Number of copies: 9 / Formula: PX4 |
|---|---|
| Molecular weight | Theoretical: 678.94 Da |
| Chemical component information | 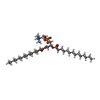 ChemComp-PX4: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 BASE (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 26.58 Å Applied symmetry - Helical parameters - Δ&Phi: -50.00 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.23 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 477012 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































