+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of lipase/ligand complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | lipase / HYDROLASE | |||||||||
| Function / homology | :  Function and homology information Function and homology information | |||||||||
| Biological species |  Acinetobacter baumannii (bacteria) Acinetobacter baumannii (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Lu DF / Zhu DY / Liu S | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2026 Journal: Cell Rep / Year: 2026Title: Structural mechanism of 3'3'-cGAMP-induced filamentation and phospholipid hydrolysis by CapV in bacterial antiphage defense. Authors: Yun Lv / Sheng Liu / Qihai Wang / Jing Zhu / Yingxiang Hou / Haiyan Xu / Deyan Zhu / Yu Liu / Jing Wu / Changxin Wu / Guijun Shang / Hongxiang Lou / Defen Lu / Huiqing Yuan / Deyu Zhu /  Abstract: The cyclic-oligonucleotide-based antiphage signaling system (CBASS) protects bacteria from phage infection. In Vibrio cholerae, phage infection activates CD-NTase DncV to produce 3'3'-cGAMP, which ...The cyclic-oligonucleotide-based antiphage signaling system (CBASS) protects bacteria from phage infection. In Vibrio cholerae, phage infection activates CD-NTase DncV to produce 3'3'-cGAMP, which triggers phospholipase CapV to degrade phosphatidylethanolamine and phosphatidylglycerol, the major phospholipids in the inner-membranes, thereby inducing cell death. However, how 3'3'-cGAMP activates CapV was unclear. Here we present crystal structures of inactive Acinetobacter baumannii CapV in apo and 3'3'-cGAMP-bound forms, along with cryo-EM structures of activated CapV-3'3'-cGAMP complex, with or without substrate dioleoylphosphatidyl-ethanolamine (DOPE). Apo-CapV forms symmetric dimers in a "closed" state. 3'3'-cGAMP binding drives lateral polymerization of dimers into filament assembly, inducing an "open" state that exposes the active site and substrate-binding cleft. DOPE binding further shifts CapV to an "ajar" state, where a Y-shaped cleft positions DOPE for hydrolysis via a conserved Ser/Asp catalytic dyad. This 3'3'-cGAMP-induced filamentation mirrors activation mechanisms of TIR-STING, TIR-SAVED, and mammalian STING, revealing a conserved signaling pattern across immune systems. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_62334.map.gz emd_62334.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-62334-v30.xml emd-62334-v30.xml emd-62334.xml emd-62334.xml | 18.5 KB 18.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_62334.png emd_62334.png | 39 KB | ||
| Filedesc metadata |  emd-62334.cif.gz emd-62334.cif.gz | 6 KB | ||
| Others |  emd_62334_half_map_1.map.gz emd_62334_half_map_1.map.gz emd_62334_half_map_2.map.gz emd_62334_half_map_2.map.gz | 58.6 MB 58.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-62334 http://ftp.pdbj.org/pub/emdb/structures/EMD-62334 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-62334 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-62334 | HTTPS FTP |
-Related structure data
| Related structure data |  9kh7MC  9kh6C  9wn0C  9wn1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_62334.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_62334.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_62334_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_62334_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : complex of lipase and ligand
| Entire | Name: complex of lipase and ligand |
|---|---|
| Components |
|
-Supramolecule #1: complex of lipase and ligand
| Supramolecule | Name: complex of lipase and ligand / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Acinetobacter baumannii (bacteria) Acinetobacter baumannii (bacteria) |
-Macromolecule #1: CBASS cGAMP-activated phospholipase
| Macromolecule | Name: CBASS cGAMP-activated phospholipase / type: protein_or_peptide / ID: 1 Details: Sequence reference for strain 'Acinetobacter baumannii' is not available in UniProt at the time of biocuration. Current sequence reference is from UniProt id A0A9E9MHI8. Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acinetobacter baumannii (bacteria) Acinetobacter baumannii (bacteria) |
| Molecular weight | Theoretical: 36.897059 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEKFQILALS GGGYRGLFTA TVLKELEQEA KENGHDSIAD CFDLITGTSV GGIVALAIAY GIKVEAIVDL FKSHGDKIFQ PKPFLKFTG SKYSNESLKT VLEEWFGDSI LGDLKCPVVI PTIDFTRGSP VTLKTPHNPN LKRDWKLKIV DVALATSAAP T YFPRHPIG ...String: MEKFQILALS GGGYRGLFTA TVLKELEQEA KENGHDSIAD CFDLITGTSV GGIVALAIAY GIKVEAIVDL FKSHGDKIFQ PKPFLKFTG SKYSNESLKT VLEEWFGDSI LGDLKCPVVI PTIDFTRGSP VTLKTPHNPN LKRDWKLKIV DVALATSAAP T YFPRHPIG PNEYVAGGLF ANDPSLIGLH EADYMFKKNI QDVHILSIGT LSSKKQLNPS TKKDGGYLDW GEGSILKAAP NI IDLVLSS QQQFMEQMVK HRMEPFPNQF YKIDEQIVQA SAQFIGLDET SDAAKQVLEG NGIQSAKVAL GKDFIRNYFN QPS RKREWF DGPQKNV UniProtKB: UNIPROTKB: A0A9E9MHI8 |
-Macromolecule #2: 2-amino-9-[(2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-9-(6-amino-9H-...
| Macromolecule | Name: 2-amino-9-[(2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-9-(6-amino-9H-purin-9-yl)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecin-2- ...Name: 2-amino-9-[(2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-9-(6-amino-9H-purin-9-yl)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecin-2-yl]-1,9-dihydro-6H-purin-6-one type: ligand / ID: 2 / Number of copies: 6 / Formula: 4BW |
|---|---|
| Molecular weight | Theoretical: 674.411 Da |
| Chemical component information | 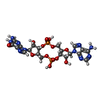 ChemComp-4BW: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 1.39 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































