[English] 日本語
 Yorodumi
Yorodumi- EMDB-6164: Detailed Structural and biochemical characterization of the nexin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6164 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Detailed Structural and biochemical characterization of the nexin-dynein regulatory complex | |||||||||
 Map data Map data | Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cilia and flagella / axoneme / dynein regulatory complex | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 45.0 Å | |||||||||
 Authors Authors | Oda T / Yanagisawa H / Kikkawa M | |||||||||
 Citation Citation |  Journal: Mol Biol Cell / Year: 2015 Journal: Mol Biol Cell / Year: 2015Title: Detailed structural and biochemical characterization of the nexin-dynein regulatory complex. Authors: Toshiyuki Oda / Haruaki Yanagisawa / Masahide Kikkawa /  Abstract: The nexin-dynein regulatory complex (N-DRC) forms a cross-bridge between the outer doublet microtubules of the axoneme and regulates dynein motor activity in cilia/flagella. Although the molecular ...The nexin-dynein regulatory complex (N-DRC) forms a cross-bridge between the outer doublet microtubules of the axoneme and regulates dynein motor activity in cilia/flagella. Although the molecular composition and the three-dimensional structure of N-DRC have been studied using mutant strains lacking N-DRC subunits, more accurate approaches are necessary to characterize the structure and function of N-DRC. In this study, we precisely localized DRC1, DRC2, and DRC4 using cryo-electron tomography and structural labeling. All three N-DRC subunits had elongated conformations and spanned the length of N-DRC. Furthermore, we purified N-DRC and characterized its microtubule-binding properties. Purified N-DRC bound to the microtubule and partially inhibited microtubule sliding driven by the outer dynein arms (ODAs). Of interest, microtubule sliding was observed even in the presence of fourfold molar excess of N-DRC relative to ODA. These results provide insights into the role of N-DRC in generating the beating motions of cilia/flagella. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6164.map.gz emd_6164.map.gz | 2.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6164-v30.xml emd-6164-v30.xml emd-6164.xml emd-6164.xml | 8.6 KB 8.6 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6164.gif 400_6164.gif 80_6164.gif 80_6164.gif | 37.4 KB 3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6164 http://ftp.pdbj.org/pub/emdb/structures/EMD-6164 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6164 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6164 | HTTPS FTP |
-Related structure data
| Related structure data |  6154C  6155C  6156C  6157C  6158C  6159C  6160C  6161C  6162C  6163C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6164.map.gz / Format: CCP4 / Size: 3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6164.map.gz / Format: CCP4 / Size: 3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
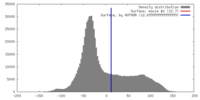
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme
| Entire | Name: Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme |
|---|---|
| Components |
|
-Supramolecule #1000: Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme
| Supramolecule | Name: Streptavidin-labeled N-DRC structure from DRC5-C-BCCP axoneme type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: axoneme
| Supramolecule | Name: axoneme / type: organelle_or_cellular_component / ID: 1 / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Concentration | 0.01 mg/mL |
|---|---|
| Buffer | pH: 7.2 Details: 30 mM Hepes-NaOH pH 7.2, 5 mM MgCl2, 1 mM dithiothreitol, 1 mM EGTA, 50 mM NaCl |
| Grid | Details: 300 mesh copper grid, holey carbon |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 98 % / Chamber temperature: 93 K / Instrument: LEICA EM GP / Method: Blot for 5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3100FFC |
|---|---|
| Specialist optics | Energy filter - Name: Omega / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Jan 3, 2014 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Average electron dose: 100 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.2 mm / Nominal defocus max: 7.0 µm / Nominal defocus min: 6.0 µm / Nominal magnification: 25700 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -65 ° / Tilt series - Axis1 - Max angle: 65 ° |
- Image processing
Image processing
| Details | Number of tilts (projections) used in 3D reconstruction: 65 Tomographic tilt angle increment: 2 |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 45.0 Å / Resolution method: OTHER / Software - Name: IMOD, PEET |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)