[English] 日本語
 Yorodumi
Yorodumi- EMDB-5767: Electron cryo-microscopy of a chemical cross-linked K87C/S199C Mm... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5767 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
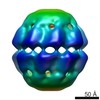
| Title | Electron cryo-microscopy of a chemical cross-linked K87C/S199C Mm-cpn with ABDM | |||||||||
 Map data Map data | mutant K87C/S199C Mm-cpn | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | protein engineering / light-gated nanocage / ABDM cross-linked Mm-cpn | |||||||||
| Biological species |  Methanococcus maripaludis (archaea) Methanococcus maripaludis (archaea) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 15.6 Å | |||||||||
 Authors Authors | Hoersch D / Roh SH / Chiu W / Kortemme T | |||||||||
 Citation Citation |  Journal: Nat Nanotechnol / Year: 2013 Journal: Nat Nanotechnol / Year: 2013Title: Reprogramming an ATP-driven protein machine into a light-gated nanocage. Authors: Daniel Hoersch / Soung-Hun Roh / Wah Chiu / Tanja Kortemme /  Abstract: Natural protein assemblies have many sophisticated architectures and functions, creating nanoscale storage containers, motors and pumps. Inspired by these systems, protein monomers have been ...Natural protein assemblies have many sophisticated architectures and functions, creating nanoscale storage containers, motors and pumps. Inspired by these systems, protein monomers have been engineered to self-assemble into supramolecular architectures including symmetrical, metal-templated and cage-like structures. The complexity of protein machines, however, has made it difficult to create assemblies with both defined structures and controllable functions. Here we report protein assemblies that have been engineered to function as light-controlled nanocontainers. We show that an adenosine-5'-triphosphate-driven group II chaperonin, which resembles a barrel with a built-in lid, can be reprogrammed to open and close on illumination with different wavelengths of light. By engineering photoswitchable azobenzene-based molecules into the structure, light-triggered changes in interatomic distances in the azobenzene moiety are able to drive large-scale conformational changes of the protein assembly. The different states of the assembly can be visualized with single-particle cryo-electron microscopy, and the nanocages can be used to capture and release non-native cargos. Similar strategies that switch atomic distances with light could be used to build other controllable nanoscale machines. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5767.map.gz emd_5767.map.gz | 48.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5767-v30.xml emd-5767-v30.xml emd-5767.xml emd-5767.xml | 8.9 KB 8.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5767_1.jpg emd_5767_1.jpg | 67.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5767 http://ftp.pdbj.org/pub/emdb/structures/EMD-5767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5767 | HTTPS FTP |
-Validation report
| Summary document |  emd_5767_validation.pdf.gz emd_5767_validation.pdf.gz | 78.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5767_full_validation.pdf.gz emd_5767_full_validation.pdf.gz | 77.5 KB | Display | |
| Data in XML |  emd_5767_validation.xml.gz emd_5767_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5767 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5767 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5767.map.gz / Format: CCP4 / Size: 51.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5767.map.gz / Format: CCP4 / Size: 51.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mutant K87C/S199C Mm-cpn | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.81 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : apo K87C/S199C mutant Mm-cpn
| Entire | Name: apo K87C/S199C mutant Mm-cpn |
|---|---|
| Components |
|
-Supramolecule #1000: apo K87C/S199C mutant Mm-cpn
| Supramolecule | Name: apo K87C/S199C mutant Mm-cpn / type: sample / ID: 1000 / Oligomeric state: homohexadecamer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 1 MDa / Theoretical: 1 MDa / Method: size exclusion column |
-Macromolecule #1: mutant Methanococcus maripaludis chaperonin
| Macromolecule | Name: mutant Methanococcus maripaludis chaperonin / type: protein_or_peptide / ID: 1 / Name.synonym: mutant Mm-cpn / Oligomeric state: 16-mer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Methanococcus maripaludis (archaea) Methanococcus maripaludis (archaea) |
| Molecular weight | Experimental: 1 MDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 50 mM KCl, 5 mM MgCl2, 20 mM HEPES, 10% glycerol, 0.1% OG |
| Grid | Details: 200-mesh R1.2/1.3 holey-carbon grids, glow discharged |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Min: 100 K |
| Date | Nov 26, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 45 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 69000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder: Gatan 626 single tilt cryo transfer holder (70) Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | EMAN2 standard package |
|---|---|
| CTF correction | Details: particle based |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 15.6 Å / Resolution method: OTHER / Software - Name: EMAN2 / Number images used: 7389 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















