[English] 日本語
 Yorodumi
Yorodumi- EMDB-5553: Molecular structure of the native HIV-1 Env trimer bound to m36: ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5553 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
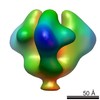
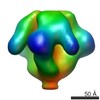
| Title | Molecular structure of the native HIV-1 Env trimer bound to m36: Membrane region | |||||||||
 Map data Map data | Molecular structure of native HIV-1 Env trimer bound to m36: Membrane regions | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | gp120 / gp41 / cryoelectron microscopy / AIDS vaccine / virus entry | |||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | subtomogram averaging / cryo EM | |||||||||
 Authors Authors | Meyerson JR / Tran EEH / Kuybeda O / Chen W / Dimitrov DS / Gorlani A / Verrips T / Lifson JD / Subramaniam S | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2013 Journal: Proc Natl Acad Sci U S A / Year: 2013Title: Molecular structures of trimeric HIV-1 Env in complex with small antibody derivatives. Authors: Joel R Meyerson / Erin E H Tran / Oleg Kuybeda / Weizao Chen / Dimiter S Dimitrov / Andrea Gorlani / Theo Verrips / Jeffrey D Lifson / Sriram Subramaniam /  Abstract: The extensive carbohydrate coat, the variability of protein structural features on HIV-1 envelope glycoproteins (Env), and the steric constraints of the virus-cell interface during infection, present ...The extensive carbohydrate coat, the variability of protein structural features on HIV-1 envelope glycoproteins (Env), and the steric constraints of the virus-cell interface during infection, present challenges to the elicitation of effective full-length (~150 kDa), neutralizing antibodies against HIV. These hurdles have motivated the engineering of smaller antibody derivatives that can bind Env and neutralize the virus. To further understand the mechanisms by which these proteins neutralize HIV-1, we carried out cryoelectron tomography of native HIV-1 BaL virions complexed separately to two small (~15 kDa) HIV-neutralizing proteins: A12, which binds the CD4-binding site on Env, and m36, whose binding to Env is enhanced by CD4 binding. We show that despite their small size, the presence of these proteins and their effects on the quaternary conformation of trimeric Env can be visualized in molecular structures derived by cryoelectron tomography combined with subvolume averaging. Binding of Env to A12 results in a conformational change that is comparable to changes observed upon its binding to the CD4-binding site antibody, b12. In contrast, binding of Env to m36 results in an "open" quaternary conformation similar to that seen with binding of soluble CD4 or the CD4i antibody, 17b. Because these small neutralizing proteins are less sterically hindered than full-length antibodies at zones of virus-cell contact, the finding that their binding has the same structural consequences as that of other broadly neutralizing antibodies highlights their potential for use in therapeutic applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5553.map.gz emd_5553.map.gz | 850 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5553-v30.xml emd-5553-v30.xml emd-5553.xml emd-5553.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5553_1.png emd_5553_1.png | 51.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5553 http://ftp.pdbj.org/pub/emdb/structures/EMD-5553 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5553 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5553 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5553.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5553.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Molecular structure of native HIV-1 Env trimer bound to m36: Membrane regions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : m36 in complex with HIV-1 Bal Env
| Entire | Name: m36 in complex with HIV-1 Bal Env |
|---|---|
| Components |
|
-Supramolecule #1000: m36 in complex with HIV-1 Bal Env
| Supramolecule | Name: m36 in complex with HIV-1 Bal Env / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Macromolecule #1: HIV-1 envelope glycoprotein
| Macromolecule | Name: HIV-1 envelope glycoprotein / type: protein_or_peptide / ID: 1 / Name.synonym: HIV-1 surface spike / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
- Sample preparation
Sample preparation
| Buffer | Details: HIV-1 BaL virions in TNE buffer treated with 1 mM Aldrithiol-2, 16mg/mL m36 in PBS buffer, pH 7.4 |
|---|---|
| Grid | Details: Quantifoil Multi-A, 200 mesh |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Specialist optics | Energy filter - Name: Gatan / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Jul 11, 2010 |
| Image recording | Category: CCD / Film or detector model: GENERIC CCD / Average electron dose: 100 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 2.5 µm / Nominal magnification: 34000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Average number of tilts used in the 3D reconstructions: 61. Average tomographic tilt angle increment: 2. |
|---|---|
| Final reconstruction | Software - Name:  IMOD IMOD |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















