+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5485 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bovine rotavirus DLP | |||||||||
 Map data Map data | Icosahedral reconstruction of rotavirus double-layered particle using the DE-12 direct electron detector without movie frame alignment | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | virus / assembly intermediate | |||||||||
| Biological species |  Bovine rotavirus Bovine rotavirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Campbell MG / Cheng A / Brilot AF / Moeller A / Lyumkis D / Veesler D / Pan J / Harrison SC / Potter CS / Carragher B / Grigorieff N | |||||||||
 Citation Citation |  Journal: Structure / Year: 2012 Journal: Structure / Year: 2012Title: Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Authors: Melody G Campbell / Anchi Cheng / Axel F Brilot / Arne Moeller / Dmitry Lyumkis / David Veesler / Junhua Pan / Stephen C Harrison / Clinton S Potter / Bridget Carragher / Nikolaus Grigorieff /  Abstract: Low-dose images obtained by electron cryo-microscopy (cryo-EM) are often affected by blurring caused by sample motion during electron beam exposure, degrading signal especially at high resolution. We ...Low-dose images obtained by electron cryo-microscopy (cryo-EM) are often affected by blurring caused by sample motion during electron beam exposure, degrading signal especially at high resolution. We show here that we can align frames of movies, recorded with a direct electron detector during beam exposure of rotavirus double-layered particles, thereby greatly reducing image blurring caused by beam-induced motion and sample stage instabilities. This procedure increases the efficiency of cryo-EM imaging and enhances the resolution obtained in three-dimensional reconstructions of the particle. Using movies in this way is generally applicable to all cryo-EM samples and should also improve the performance of midrange electron microscopes that may have limited mechanical stability and beam coherence. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5485.map.gz emd_5485.map.gz | 202.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5485-v30.xml emd-5485-v30.xml emd-5485.xml emd-5485.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5485_1.png emd_5485_1.png | 352.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5485 http://ftp.pdbj.org/pub/emdb/structures/EMD-5485 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5485 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5485 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5485.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5485.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral reconstruction of rotavirus double-layered particle using the DE-12 direct electron detector without movie frame alignment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.424 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
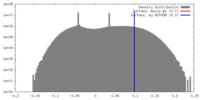
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Rotavirus double-layered particle
| Entire | Name: Rotavirus double-layered particle |
|---|---|
| Components |
|
-Supramolecule #1000: Rotavirus double-layered particle
| Supramolecule | Name: Rotavirus double-layered particle / type: sample / ID: 1000 / Details: The sample was monodisperse. Oligomeric state: 12 x VP1, 120 x VP2, 12 x VP3, 780 x VP6, 11 x RNA Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 70 MDa |
-Supramolecule #1: Bovine rotavirus
| Supramolecule | Name: Bovine rotavirus / type: virus / ID: 1 / NCBI-ID: 10927 / Sci species name: Bovine rotavirus / Database: NCBI / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Theoretical: 70 MDa |
| Virus shell | Shell ID: 1 / Name: DLP / Diameter: 700 Å / T number (triangulation number): 13 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Details: 1.2-1.3 C-flat |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK II / Method: Blot for 7 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Nov 28, 2011 |
| Image recording | Category: CCD / Film or detector model: DIRECT ELECTRON DE-12 (4k x 3k) / Digitization - Sampling interval: 6 µm / Number real images: 501 / Average electron dose: 32 e/Å2 / Details: Movies recorded at 25 frames/second |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 42135 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Manual particle selection, refinement using Frealign. |
|---|---|
| CTF correction | Details: Each particle |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.5 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: Frealign Details: Final map was calculated from 807 particles with icosahedral symmetry imposed Number images used: 807 |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)