+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human CAK bound to OTS964 | |||||||||
 Map data Map data | Post-processed (sharpened, filtered) cryo-EM map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | kinase / cyclin-dependent kinase / inhibitor / CELL CYCLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationRNA polymerase II CTD heptapeptide repeat S5 kinase activity / ventricular system development / snRNA transcription by RNA polymerase II / transcription factor TFIIK complex / CAK-ERCC2 complex / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / adult heart development / [RNA-polymerase]-subunit kinase ...RNA polymerase II CTD heptapeptide repeat S5 kinase activity / ventricular system development / snRNA transcription by RNA polymerase II / transcription factor TFIIK complex / CAK-ERCC2 complex / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / adult heart development / [RNA-polymerase]-subunit kinase / RNA Polymerase I Transcription Termination / cyclin-dependent protein serine/threonine kinase regulator activity / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Polymerase I Transcription Initiation / RNA polymerase II transcribes snRNA genes / regulation of G1/S transition of mitotic cell cycle / cyclin-dependent kinase / cyclin-dependent protein serine/threonine kinase activity / ATP-dependent activity, acting on DNA / Tat-mediated elongation of the HIV-1 transcript / Cyclin E associated events during G1/S transition / Formation of HIV-1 elongation complex containing HIV-1 Tat / Cyclin A:Cdk2-associated events at S phase entry / Formation of HIV elongation complex in the absence of HIV Tat / cyclin-dependent protein kinase holoenzyme complex / Cyclin A/B1/B2 associated events during G2/M transition / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / RNA polymerase II CTD heptapeptide repeat kinase activity / RNA Polymerase II Pre-transcription Events / positive regulation of smooth muscle cell proliferation / male germ cell nucleus / TP53 Regulates Transcription of DNA Repair Genes / nucleotide-excision repair / transcription initiation at RNA polymerase II promoter / RNA Polymerase I Promoter Escape / G1/S transition of mitotic cell cycle / NoRC negatively regulates rRNA expression / response to calcium ion / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex / fibrillar center / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / Cyclin D associated events in G1 / kinase activity / Gap-filling DNA repair synthesis and ligation in TC-NER / RUNX1 regulates transcription of genes involved in differentiation of HSCs / transcription by RNA polymerase II / protein kinase activity / regulation of cell cycle / protein stabilization / cell division / protein serine kinase activity / DNA repair / protein serine/threonine kinase activity / regulation of transcription by RNA polymerase II / negative regulation of apoptotic process / perinuclear region of cytoplasm / positive regulation of transcription by RNA polymerase II / zinc ion binding / nucleoplasm / ATP binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.4 Å | |||||||||
 Authors Authors | McGeoch AJS / Cushing VI / Cronin NB / Alfieri C / Greber BJ | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2026 Journal: Nat Commun / Year: 2026Title: Cryo-EM structures of the CDK11-cyclin L-SAP30BP complex reveal mechanisms of CDK11 regulation. Authors: Amy J S McGeoch / Victoria I Cushing / Theodoros I Roumeliotis / Nora B Cronin / Stephen J Hearnshaw / Jyoti S Choudhary / Claudio Alfieri / Basil J Greber /  Abstract: The cyclin-dependent kinase CDK11 functions in transcription, mitotic progression, and mRNA splicing. Specifically, spliceosome activation during the B to B transition depends on phosphorylation of ...The cyclin-dependent kinase CDK11 functions in transcription, mitotic progression, and mRNA splicing. Specifically, spliceosome activation during the B to B transition depends on phosphorylation of the U2 snRNP component SF3B1 by the CDK11-cyclin L-SAP30BP complex. Here, we present the structure of this spliceosome-activating CDK-cyclin complex, determined by cryogenic electron microscopy at 2.3 Å resolution. Our structure and biochemical experiments show that SAP30BP forms extensive interactions with cyclin L2, thereby stabilising it, and forms critical interactions with the C-terminal kinase lobe of CDK11 that promote complex assembly. Destabilisation of cyclin L2 in the absence of SAP30BP suggests that these principles are applicable to all CDK11-cyclin L complexes. Furthermore, we identify a pseudo-substrate sequence near the CDK11 C-terminus and provide evidence for a role of this segment in CDK11 auto-regulation. Finally, the structure of the CDK11-cyclin L-SAP30BP complex bound to the clinical high-affinity CDK11 inhibitor OTS964 and a comparison to OTS964-bound off-target complexes provide insight into the mechanism of OTS964 selectivity and specificity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_53205.map.gz emd_53205.map.gz | 25.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-53205-v30.xml emd-53205-v30.xml emd-53205.xml emd-53205.xml | 26.3 KB 26.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_53205.png emd_53205.png | 115.2 KB | ||
| Masks |  emd_53205_msk_1.map emd_53205_msk_1.map | 27 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-53205.cif.gz emd-53205.cif.gz | 7.4 KB | ||
| Others |  emd_53205_half_map_1.map.gz emd_53205_half_map_1.map.gz emd_53205_half_map_2.map.gz emd_53205_half_map_2.map.gz | 20.7 MB 20.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-53205 http://ftp.pdbj.org/pub/emdb/structures/EMD-53205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-53205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-53205 | HTTPS FTP |
-Related structure data
| Related structure data |  9qjnMC  9qjjC  9qktC  9qkzC  9ql1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_53205.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_53205.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed (sharpened, filtered) cryo-EM map. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_53205_msk_1.map emd_53205_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unfiltered cryo-EM half map.
| File | emd_53205_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered cryo-EM half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unfiltered cryo-EM half map.
| File | emd_53205_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered cryo-EM half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CAK-OTS964 complex
| Entire | Name: CAK-OTS964 complex |
|---|---|
| Components |
|
-Supramolecule #1: CAK-OTS964 complex
| Supramolecule | Name: CAK-OTS964 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 80 KDa |
-Macromolecule #1: CDK-activating kinase assembly factor MAT1
| Macromolecule | Name: CDK-activating kinase assembly factor MAT1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.234531 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAPVTFSTG IKMGQHISLA PIHKLEEALY EYQPLQIETY GPHVPELEML GRLGYLNHVR AASPQDLAGG YTSSLACHRA LQDAFSGLF WQPS UniProtKB: CDK-activating kinase assembly factor MAT1 |
-Macromolecule #2: Cyclin-H
| Macromolecule | Name: Cyclin-H / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.721508 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: (ACE)MYHNSSQKR HWTFSSEEQL ARLRADANRK FRCKAVANGK VLPNDPVFLE PHEEMTLCKY YEKRLLEFCS VFKPAM PRS VVGTACMYFK RFYLNNSVME YHPRIIMLTC AFLACKVDEF NVSSPQFVGN LRESPLGQEK ALEQILEYEL LLIQQLN FH LIVHNPYRPF ...String: (ACE)MYHNSSQKR HWTFSSEEQL ARLRADANRK FRCKAVANGK VLPNDPVFLE PHEEMTLCKY YEKRLLEFCS VFKPAM PRS VVGTACMYFK RFYLNNSVME YHPRIIMLTC AFLACKVDEF NVSSPQFVGN LRESPLGQEK ALEQILEYEL LLIQQLN FH LIVHNPYRPF EGFLIDLKTR YPILENPEIL RKTADDFLNR IALTDAYLLY TPSQIALTAI LSSASRAGIT MESYLSES L MLKENRTCLS QLLDIMKSMR NLVKKYEPPR SEEVAVLKQK LERCHSAELA LNVITKKRKG YEDDDYVSKK SKHEEEEWT DDDLVESL UniProtKB: Cyclin-H |
-Macromolecule #3: Cyclin-dependent kinase 7
| Macromolecule | Name: Cyclin-dependent kinase 7 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: cyclin-dependent kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.362598 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAMALDVKS RAKRYEKLDF LGEGQFATVY KARDKNTNQI VAIKKIKLGH RSEAKDGINR TALREIKLLQ ELSHPNIIGL LDAFGHKSN ISLVFDFMET DLEVIIKDNS LVLTPSHIKA YMLMTLQGLE YLHQHWILHR DLKPNNLLLD ENGVLKLADF G LAKSFGSP ...String: SNAMALDVKS RAKRYEKLDF LGEGQFATVY KARDKNTNQI VAIKKIKLGH RSEAKDGINR TALREIKLLQ ELSHPNIIGL LDAFGHKSN ISLVFDFMET DLEVIIKDNS LVLTPSHIKA YMLMTLQGLE YLHQHWILHR DLKPNNLLLD ENGVLKLADF G LAKSFGSP NRAYTHQVVT RWYRAPELLF GARMYGVGVD MWAVGCILAE LLLRVPFLPG DSDLDQLTRI FETLGTPTEE QW PDMCSLP DYVTFKSFPG IPLHHIFSAA GDDLLDLIQG LFLFNPCARI TATQALKMKY FSNRPGPTPG CQLPRPNCPV ETL KEQSNP ALAIKRKRTE ALEQGGLPKK LIF UniProtKB: Cyclin-dependent kinase 7 |
-Macromolecule #4: OTS964
| Macromolecule | Name: OTS964 / type: ligand / ID: 4 / Number of copies: 1 / Formula: NK3 |
|---|---|
| Molecular weight | Theoretical: 392.514 Da |
| Chemical component information | 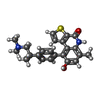 ChemComp-NK3: |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 142 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
| ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 10788 / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)














































