[English] 日本語
 Yorodumi
Yorodumi- EMDB-51885: Haemophilus influenzae serotype b capsule polymerase bcs3 in comp... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Haemophilus influenzae serotype b capsule polymerase bcs3 in complex with acceptor polymer | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Capsule polymerase / Haemophilus influenzae serotype b / Pathogens / BIOSYNTHETIC PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationCDP-glycerol glycerophosphotransferase activity / teichoic acid biosynthetic process / metal ion binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Haemophilus influenzae (bacteria) Haemophilus influenzae (bacteria) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.65 Å | ||||||||||||
 Authors Authors | Di Domenico V / Mast T / Cifuente JO / Schulze J / Litschko C / Alcaide A / Villegas JC / Fiebig T / Guerin ME | ||||||||||||
| Funding support |  Spain, Spain,  United States, United States,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Haemophilus influenzae serotype b capsule polymerase bcs3 in complex with acceptor polymer Authors: Di Domenico V / Mast T / Cifuente JO / Schulze J / Litschko C / Alcaide A / Villegas JC / Fiebig T / Guerin ME | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_51885.map.gz emd_51885.map.gz | 88.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-51885-v30.xml emd-51885-v30.xml emd-51885.xml emd-51885.xml | 22.5 KB 22.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_51885_fsc.xml emd_51885_fsc.xml | 16.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_51885.png emd_51885.png | 83.1 KB | ||
| Filedesc metadata |  emd-51885.cif.gz emd-51885.cif.gz | 7 KB | ||
| Others |  emd_51885_additional_1.map.gz emd_51885_additional_1.map.gz emd_51885_half_map_1.map.gz emd_51885_half_map_1.map.gz emd_51885_half_map_2.map.gz emd_51885_half_map_2.map.gz | 158.2 MB 164.9 MB 164.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-51885 http://ftp.pdbj.org/pub/emdb/structures/EMD-51885 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51885 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51885 | HTTPS FTP |
-Related structure data
| Related structure data |  9h5iMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_51885.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_51885.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.653 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Sharpened map
| File | emd_51885_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_51885_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_51885_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Physiological dimeric assembly of Bcs3
| Entire | Name: Physiological dimeric assembly of Bcs3 |
|---|---|
| Components |
|
-Supramolecule #1: Physiological dimeric assembly of Bcs3
| Supramolecule | Name: Physiological dimeric assembly of Bcs3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Haemophilus influenzae (bacteria) Haemophilus influenzae (bacteria) |
-Macromolecule #1: Bcs3
| Macromolecule | Name: Bcs3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haemophilus influenzae (bacteria) Haemophilus influenzae (bacteria) |
| Molecular weight | Theoretical: 137.381172 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VDKTWLFGSY AWQGNPKALF LYMLVNCKET HECWWVADNE ESMKSIKKST GLKNITFTDS EKAKELFPHA DVYVTENFRE SYPVYMNEN IKVFNTWHGV GLKHIELALG MNSVLAESIV RKYVRNYDIY KNNVLFLTTS QAMEDHFLED MAISKELIIR G KYPRNAVY ...String: VDKTWLFGSY AWQGNPKALF LYMLVNCKET HECWWVADNE ESMKSIKKST GLKNITFTDS EKAKELFPHA DVYVTENFRE SYPVYMNEN IKVFNTWHGV GLKHIELALG MNSVLAESIV RKYVRNYDIY KNNVLFLTTS QAMEDHFLED MAISKELIIR G KYPRNAVY GPNGIHTYDI NTLLPKNKSQ YSQTILFCPT YRIGAIQGVL NSLLPDFAKL EEVCRHKNQL FIVKVHPFMK KD NYFAEMS EKYKDSEYIL FWNDDYDIYE AFNSIDLAII DYSSIFYDLL DAGVEKFIRY VPDLDEYQND LELIGDYADL TEG RIVKSF QQLLNCLDNA NIKIISTKRK QYLMDYFFGF KKENKSMESL IADVDNCQLQ PKSLKELHTF DIFDTLIRRS TLRP FSIFD YVRDKAKASG IKFPLALTEN WINVRNRAEH DVRDIMRKTT FERQSDKIEI TLDDIYTRLQ KNLLLTDEQT DFLKQ AEIE AEIAHVEPIQ KRINYLFSLK AKGHDVAMAS DMYLPEDVIY KMLDRADTRL REIPLYLSST IGYQKSTGKL YQHIFF DLD YQYSRWTHYG DNKHADGSVP RRLGIQTAVH DIDDFIPFEN AMVNAMDNYN RYPAYQLATK MHRYRTQLVQ ENGFGNT LF ETKYYNYAYV GASFVPYINW AIKDAIKRGY ETIYFISRDG HFLKQIADKI IEIRGYNVKT KYIYGSRKAW RLPSFITK V DDETFWQFGN FVGMDSFEDL VKASYLSESE LLSLFPEFES LRHAKHLRGE IAENIRKIFK NSPAYHEKVL AIAAEKRKM VRQYIQQEIN PKEKFAFVEF WGRGYTQDTF GRLLNDAFGK EVKNPFYYVR SFTDDMGTSV RHNFILAPQN FSFFEPIFAQ TPYDSIPDY YEEKGRIEPI INHRDRSVSD LISEGLLKFT EDYLALNTQD EDYFDAALSQ FNYQYQLNTP NDQFICNVFS E LKDNISSF GVEKPYAPAL TLKQLESITS KQELDKLTQS IPISLSKSDV KVIDYYNKIQ KNYNLPAYNS TPMRKAYAVN PL EQYVWST QVPFRVLSLK QNSFYLDVSF AETTKRKDIF LKELNEIDVI AVDWLKGGVP RLLTEHGYIT AHKDWVKKSF NDD KTNNIE EPKVKNKEKS KVLEVNTTVT NNNKQAIGKL DNNIDKSLEH HHHHH UniProtKB: Bcs3 |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: beta-D-ribosyl-(1->1)-D-ribitol-5-phosphate
| Macromolecule | Name: beta-D-ribosyl-(1->1)-D-ribitol-5-phosphate / type: ligand / ID: 3 / Number of copies: 12 / Formula: KOF |
|---|---|
| Molecular weight | Theoretical: 364.24 Da |
| Chemical component information | 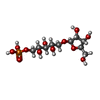 ChemComp-KOF: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 72 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 27103 / Average electron dose: 51.85 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)














































