[English] 日本語
 Yorodumi
Yorodumi- EMDB-51410: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-8... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-818) dimer in the empty conformation | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mycoplasma pneumoniae Lipid Transfer Lipid Transport / LIPID BINDING PROTEIN | |||||||||
| Biological species |  Mycoplasmoides pneumoniae M129 (bacteria) Mycoplasmoides pneumoniae M129 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.52 Å | |||||||||
 Authors Authors | Manger S | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Essential protein P116 extracts cholesterol and other indispensable lipids for Mycoplasmas. Authors: Lasse Sprankel / David Vizarraga / Jesús Martín / Sina Manger / Jakob Meier-Credo / Marina Marcos / Josep Julve / Noemi Rotllan / Margot P Scheffer / Joan Carles Escolà-Gil / Julian D ...Authors: Lasse Sprankel / David Vizarraga / Jesús Martín / Sina Manger / Jakob Meier-Credo / Marina Marcos / Josep Julve / Noemi Rotllan / Margot P Scheffer / Joan Carles Escolà-Gil / Julian D Langer / Jaume Piñol / Ignacio Fita / Achilleas S Frangakis /   Abstract: Mycoplasma pneumoniae, responsible for approximately 30% of community-acquired human pneumonia, needs to extract lipids from the host environment for survival and proliferation. Here, we report a ...Mycoplasma pneumoniae, responsible for approximately 30% of community-acquired human pneumonia, needs to extract lipids from the host environment for survival and proliferation. Here, we report a comprehensive structural and functional analysis of the previously uncharacterized protein P116 (MPN_213). Single-particle cryo-electron microscopy of P116 reveals a homodimer presenting a previously unseen fold, forming a huge hydrophobic cavity, which is fully accessible to solvent. Lipidomics analysis shows that P116 specifically extracts lipids such as phosphatidylcholine, sphingomyelin and cholesterol. Structures of different conformational states reveal the mechanism by which lipids are extracted. This finding immediately suggests a way to control Mycoplasma infection by interfering with lipid uptake. #1:  Journal: Biorxiv / Year: 2024 Journal: Biorxiv / Year: 2024Title: A Self-Sufficient Lipid Transport Protein in Mycoplasma pneumoniae Authors: Manger S / Arghittu SM / Sprankel L / Meier-Credo J / Wieland K / Bublak D / Langer J / Covino R / Frangakis AS | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_51410.map.gz emd_51410.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-51410-v30.xml emd-51410-v30.xml emd-51410.xml emd-51410.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_51410.png emd_51410.png | 55.6 KB | ||
| Masks |  emd_51410_msk_1.map emd_51410_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-51410.cif.gz emd-51410.cif.gz | 5.7 KB | ||
| Others |  emd_51410_half_map_1.map.gz emd_51410_half_map_1.map.gz emd_51410_half_map_2.map.gz emd_51410_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-51410 http://ftp.pdbj.org/pub/emdb/structures/EMD-51410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-51410 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_51410.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_51410.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.674 Å | ||||||||||||||||||||||||||||||||||||
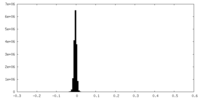
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_51410_msk_1.map emd_51410_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
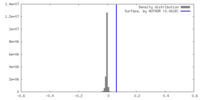
| Density Histograms |
-Half map: #1
| File | emd_51410_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
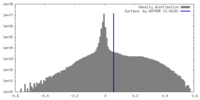
| Density Histograms |
-Half map: #2
| File | emd_51410_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
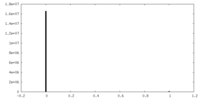
| Density Histograms |
- Sample components
Sample components
-Entire : Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-8...
| Entire | Name: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-818) dimer in the empty conformation |
|---|---|
| Components |
|
-Supramolecule #1: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-8...
| Supramolecule | Name: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-818) dimer in the empty conformation type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Mycoplasmoides pneumoniae M129 (bacteria) Mycoplasmoides pneumoniae M129 (bacteria) |
-Macromolecule #1: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-8...
| Macromolecule | Name: Mycoplasma pneumoniae protein P116 truncated ectodomain (aa 246-818) dimer in the empty conformation type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycoplasmoides pneumoniae M129 (bacteria) Mycoplasmoides pneumoniae M129 (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: GVDVFEAQKN LVGKGKYLNT HVKAEDVKKD VNANIKNQFD IAKIIAELMG KALKEFGNQQ EGQPLSFLK VMDKVKEDFE KLFNLVRPGL GKFVKDLIQS SSQAENKITV YKLIFDNKKT I LNLLKELS IPELNSSLGL VDVLFDGITD SDGLYERLQS FKDLIVPAVK ...String: GVDVFEAQKN LVGKGKYLNT HVKAEDVKKD VNANIKNQFD IAKIIAELMG KALKEFGNQQ EGQPLSFLK VMDKVKEDFE KLFNLVRPGL GKFVKDLIQS SSQAENKITV YKLIFDNKKT I LNLLKELS IPELNSSLGL VDVLFDGITD SDGLYERLQS FKDLIVPAVK TNEKTAALSP LI EELLTQK DTYVFDLIQK HKGILTNLLK NFLADFQKST PFMADQVAIF TELFDNEGAF DLF GEADFV DKIAELFLTK RTVKNGEKIE TKDSLLVTSL KSLLGEKVAA LGDLLDSYIF KNEL LNRSV EVAKAEAKDT KGATDYKKEQ AKALKKLFKH IGENTLSKTN LDKITLKEVK NTENV ELEE TETTLKVKKL DVEYKVELGN FEIKNGLIKA MLEFLPDTKD LETTLDKLLF KGESYKAMKD KYIKEGFPGY GWAKGVVPGA FESIENTFKS AIDKTKSIRD LFGDMLFGND LSSVKETDSF ITLGGSFDIK YGGENLNVLP AYYSLINSEI GYQIIGVDTT IDATKVKVEL KNKEYKGKSP AINGQVKLSQ SFFNVWTNMF DSITKQIFQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM Tris-HCl, 1 mM CHAPSO |
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)