[English] 日本語
 Yorodumi
Yorodumi- EMDB-5133: 17 angstrom of wild Rabbit Hemorrhagic DiseaseVirus core-like par... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5133 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 17 angstrom of wild Rabbit Hemorrhagic DiseaseVirus core-like particle cryomicroscopy structure | |||||||||
 Map data Map data | This is the cryo-electron microscopy reconstruction of the wild rabbit hemorrhagic disease virus core-like particles. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RHDV / vp60 / vp10 / core-like particles | |||||||||
| Biological species |  rabbit hemorrhagic disease virus core like particles rabbit hemorrhagic disease virus core like particles | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 17.0 Å | |||||||||
 Authors Authors | Hu Z / Tian X / Zhai Y / Xu W / Zheng D / Sun F | |||||||||
 Citation Citation |  Journal: Protein Cell / Year: 2010 Journal: Protein Cell / Year: 2010Title: Cryo-electron microscopy reconstructions of two types of wild rabbit hemorrhagic disease viruses characterized the structural features of Lagovirus. Authors: Zhongjun Hu / Xiaojuan Tian / Yujia Zhai / Wei Xu / Dong Zheng / Fei Sun /  Abstract: Rabbit hemorrhagic disease was described in China in 1984 and can cause hemorrhagic necrosis of the liver within two or three days after infection. The etiological agent, rabbit hemorrhagic disease ...Rabbit hemorrhagic disease was described in China in 1984 and can cause hemorrhagic necrosis of the liver within two or three days after infection. The etiological agent, rabbit hemorrhagic disease virus (RHDV), belongs to the Lagovirus genus in the Caliciviridae family. Compared to other calicivirus, such as rNV and SMSV, the structure of Lagovirus members is not well characterized. In this report, structures of two types of wild RHDV particles, the intact virion and the core-like particle (CLP), were reconstructed by cryo-electron microscopy at 11 &0A and 17 &0A, respectively. This is the first time the 3D structure of wild caliciviruses CLP has been provided, and the 3D structure of intact RHDV virion is the highest resolution structure in Lagovirus. Comparison of the intact virion and CLP structures clearly indicated that CLP was produced from the intact virion with the protrusion dissociated. In contrast with the crystal structures of recombinant Norovirus and San Miguel sea lion virus, the capsomers of RHDV virion exhibited unique structural features and assembly modes. Both P1 and P2 subdomains have interactions inside the AB capsomer, while only P2 subdomains have interaction inside CC capsomer. The pseudo atomic models of RHDV capsomers were constructed by homology modeling and density map fitting, and the rotation of RHDV VP60 P domain with respect to its S domain, compared with SMSV, was observed. Collectively, our cryo-electron microscopic studies of RHDV provide close insight into the structure of Lagovirus, which is important for functional analysis and better vaccine development in the future. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5133.map.gz emd_5133.map.gz | 11.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5133-v30.xml emd-5133-v30.xml emd-5133.xml emd-5133.xml | 10.2 KB 10.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5133_1.jpg emd_5133_1.jpg | 51.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5133 http://ftp.pdbj.org/pub/emdb/structures/EMD-5133 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5133 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5133 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5133.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5133.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the cryo-electron microscopy reconstruction of the wild rabbit hemorrhagic disease virus core-like particles. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.54 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
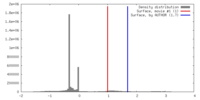
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : wild Rabbit Hemorrhagic Disease Viruses core-like particles
| Entire | Name: wild Rabbit Hemorrhagic Disease Viruses core-like particles |
|---|---|
| Components |
|
-Supramolecule #1000: wild Rabbit Hemorrhagic Disease Viruses core-like particles
| Supramolecule | Name: wild Rabbit Hemorrhagic Disease Viruses core-like particles type: sample / ID: 1000 / Oligomeric state: icosahedral / Number unique components: 3 |
|---|---|
| Molecular weight | Theoretical: 7.2 MDa |
-Supramolecule #1: rabbit hemorrhagic disease virus core like particles
| Supramolecule | Name: rabbit hemorrhagic disease virus core like particles / type: virus / ID: 1 Name.synonym: rabbit hemorrhagic disease virus core like particles Sci species name: rabbit hemorrhagic disease virus core like particles Database: NCBI / Virus type: VIROID / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No Syn species name: rabbit hemorrhagic disease virus core like particles |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Theoretical: 7.2 MDa |
| Virus shell | Shell ID: 1 / Name: S / Diameter: 320 Å / T number (triangulation number): 3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Grid | Details: 300 mesh holygrid |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER Method: Embeded in thin layer of vitreous ice on freshly carbon-coated home-made holey EM grid by manually blotting the grids with filter paper and then plunging into liquid ethane cooled by liquid nitrogen |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Temperature | Average: 105 K |
| Image recording | Category: CCD / Film or detector model: KODAK SO-163 FILM / Digitization - Sampling interval: 12.7 µm / Number real images: 230 / Average electron dose: 20 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN HELIUM |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 17.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Eman spider / Number images used: 895 |
| Final two d classification | Number classes: 21 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  chimera chimera |
| Details | Protocol: Rigid Body |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: correlation coefficient |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)