+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | mus musculus ribosome from SOLIST lamellas | |||||||||
 Map data Map data | after M refinement, filtered, sharpened | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | in situ / ribosome / mouse / mus musculus | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / Resolution: 8.3 Å | |||||||||
 Authors Authors | Erdmann PS / Nguyen HTD / Perone G / Klena N / Vazzana R / Kaluthantrige Don F / Silva M / Sorrentino S / Swuec P / Leroux F ...Erdmann PS / Nguyen HTD / Perone G / Klena N / Vazzana R / Kaluthantrige Don F / Silva M / Sorrentino S / Swuec P / Leroux F / Kalebic N / Coscia F | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2024 Journal: Nat Methods / Year: 2024Title: Serialized on-grid lift-in sectioning for tomography (SOLIST) enables a biopsy at the nanoscale. Authors: Ho Thuy Dung Nguyen / Gaia Perone / Nikolai Klena / Roberta Vazzana / Flaminia Kaluthantrige Don / Malan Silva / Simona Sorrentino / Paolo Swuec / Frederic Leroux / Nereo Kalebic / Francesca ...Authors: Ho Thuy Dung Nguyen / Gaia Perone / Nikolai Klena / Roberta Vazzana / Flaminia Kaluthantrige Don / Malan Silva / Simona Sorrentino / Paolo Swuec / Frederic Leroux / Nereo Kalebic / Francesca Coscia / Philipp S Erdmann /   Abstract: Cryo-focused ion beam milling has substantially advanced our understanding of molecular processes by opening windows into cells. However, applying this technique to complex samples, such as tissues, ...Cryo-focused ion beam milling has substantially advanced our understanding of molecular processes by opening windows into cells. However, applying this technique to complex samples, such as tissues, has presented considerable technical challenges. Here we introduce an innovative adaptation of the cryo-lift-out technique, serialized on-grid lift-in sectioning for tomography (SOLIST), addressing these limitations. SOLIST enhances throughput, minimizes ice contamination and improves sample stability for cryo-electron tomography. It thereby facilitates the high-resolution imaging of a wide range of specimens. We illustrate these advantages on reconstituted liquid-liquid phase-separated droplets, brain organoids and native tissues from the mouse brain, liver and heart. With SOLIST, cellular processes can now be investigated at molecular resolution directly in native tissue. Furthermore, our method has a throughput high enough to render cryo-lift-out a competitive tool for structural biology. This opens new avenues for unprecedented insights into cellular function and structure in health and disease, a 'biopsy at the nanoscale'. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_50630.map.gz emd_50630.map.gz | 195 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-50630-v30.xml emd-50630-v30.xml emd-50630.xml emd-50630.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_50630_fsc.xml emd_50630_fsc.xml | 13.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_50630.png emd_50630.png | 145.6 KB | ||
| Masks |  emd_50630_msk_1.map emd_50630_msk_1.map | 209.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-50630.cif.gz emd-50630.cif.gz | 4.9 KB | ||
| Others |  emd_50630_half_map_1.map.gz emd_50630_half_map_1.map.gz emd_50630_half_map_2.map.gz emd_50630_half_map_2.map.gz | 107.9 MB 107.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-50630 http://ftp.pdbj.org/pub/emdb/structures/EMD-50630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50630 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_50630.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_50630.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | after M refinement, filtered, sharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2 Å | ||||||||||||||||||||||||||||||||||||
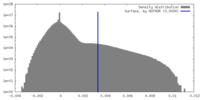
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_50630_msk_1.map emd_50630_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_50630_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_50630_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mus musculus ribosome from SOLIST lamellas
| Entire | Name: mus musculus ribosome from SOLIST lamellas |
|---|---|
| Components |
|
-Supramolecule #1: mus musculus ribosome from SOLIST lamellas
| Supramolecule | Name: mus musculus ribosome from SOLIST lamellas / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
 Processing Processing | subtomogram averaging |
|---|---|
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE Details: glow-discharged at 30 mA for 30 s (GloQube Plus Glow discharge system, Quorum Technologies) |
| Details | SOLIST lamella preparation of mouse liver from HPF. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)