[English] 日本語
 Yorodumi
Yorodumi- EMDB-4550: Cryo-EM structure of human norovirus GII.4 NSW-2012 VLP with T=4 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4550 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human norovirus GII.4 NSW-2012 VLP with T=4 icosahedral symmetry | |||||||||
 Map data Map data | Cryo-EM structure of GII.4 NSW virus like particle | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Norovirus Norovirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.3 Å | |||||||||
 Authors Authors | Devant J / Hansman G | |||||||||
 Citation Citation |  Journal: Antiviral Res / Year: 2019 Journal: Antiviral Res / Year: 2019Title: Heterologous expression of human norovirus GII.4 VP1 leads to assembly of T=4 virus-like particles. Authors: Jessica M Devant / Götz Hofhaus / David Bhella / Grant S Hansman /   Abstract: Human noroviruses are a leading cause of acute gastroenteritis, yet there are still no vaccines or antivirals available. Expression of the norovirus capsid protein (VP1) in insect cells typically ...Human noroviruses are a leading cause of acute gastroenteritis, yet there are still no vaccines or antivirals available. Expression of the norovirus capsid protein (VP1) in insect cells typically results in the formation of virus-like particles (VLPs) that are morphologically and antigenically comparable to native virions. Indeed, several different norovirus VLP candidates are currently used in clinical trials. So far, structural analysis of norovirus VLPs showed that the capsid has a T = 3 icosahedral symmetry and is composed of 180 copies of VP1 that are folded into three quasi-equivalent subunits (A, B, and C). In this study, the VLP structures of two norovirus GII.4 genetic variants that were identified in 1974 and 2012 were determined using cryo-EM. Surprisingly, we found that greater than 95% of these GII.4 VLPs were larger than virions and 3D reconstruction showed that these VLPs exhibited T = 4 icosahedral symmetry. We also discovered that the T = 4 VLPs presented several novel structural features. The T = 4 particles assembled from 240 copies of VP1 that adopted four quasi-equivalent conformations (A, B, C, and D) and formed two distinct dimers, A/B and C/D. The protruding domains were elevated ∼21 Å off the capsid shell, which was ∼7 Å more than in the previously studied GII.10 T = 3 VLPs. A small cavity and flap-like structure at the icosahedral two-fold axis disrupted the contiguous T = 4 shell. Overall, our findings indicated that GII.4 VP1 sequences assemble into T = 4 VLPs and these larger particles might have important consequences for VLP-based vaccine development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4550.map.gz emd_4550.map.gz | 58.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4550-v30.xml emd-4550-v30.xml emd-4550.xml emd-4550.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4550.png emd_4550.png | 69.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4550 http://ftp.pdbj.org/pub/emdb/structures/EMD-4550 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4550 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4550 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4550.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4550.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of GII.4 NSW virus like particle | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.27 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
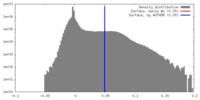
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Norovirus
| Entire | Name:  Norovirus Norovirus |
|---|---|
| Components |
|
-Supramolecule #1: Norovirus
| Supramolecule | Name: Norovirus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 142786 / Sci species name: Norovirus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Host system | Organism:  Trichoplusia ni (cabbage looper) / Recombinant strain: High 5 cells Trichoplusia ni (cabbage looper) / Recombinant strain: High 5 cells |
| Molecular weight | Theoretical: 13.4 MDa |
| Virus shell | Shell ID: 1 / Name: VP1 / Diameter: 500.0 Å / T number (triangulation number): 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Name: PBS |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER/RHODIUM / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number grids imaged: 1 / Number real images: 364 / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 64000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 7.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 2.1) / Number images used: 10548 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)