+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4510 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo em structure of the Listeria stressosome | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Listeria monocytogenes (bacteria) / Listeria monocytogenes (bacteria) /  Listeria monocytogenes EGD-E (bacteria) Listeria monocytogenes EGD-E (bacteria) | ||||||||||||
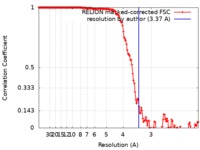
| Method | single particle reconstruction / cryo EM / Resolution: 3.37 Å | ||||||||||||
 Authors Authors | Williams AH / Redzej A | ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: The cryo-electron microscopy supramolecular structure of the bacterial stressosome unveils its mechanism of activation. Authors: Allison H Williams / Adam Redzej / Nathalie Rolhion / Tiago R D Costa / Aline Rifflet / Gabriel Waksman / Pascale Cossart /   Abstract: How the stressosome, the epicenter of the stress response in bacteria, transmits stress signals from the environment has remained elusive. The stressosome consists of multiple copies of three ...How the stressosome, the epicenter of the stress response in bacteria, transmits stress signals from the environment has remained elusive. The stressosome consists of multiple copies of three proteins RsbR, RsbS and RsbT, a kinase that is important for its activation. Using cryo-electron microscopy, we determined the atomic organization of the Listeria monocytogenes stressosome at 3.38 Å resolution. RsbR and RsbS are organized in a 60-protomers truncated icosahedron. A key phosphorylation site on RsbR (T209) is partially hidden by an RsbR flexible loop, whose "open" or "closed" position could modulate stressosome activity. Interaction between three glutamic acids in the N-terminal domain of RsbR and the membrane-bound mini-protein Prli42 is essential for Listeria survival to stress. Together, our data provide the atomic model of the stressosome core and highlight a loop important for stressosome activation, paving the way towards elucidating the mechanism of signal transduction by the stressosome in bacteria. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4510.map.gz emd_4510.map.gz | 22.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4510-v30.xml emd-4510-v30.xml emd-4510.xml emd-4510.xml | 10.6 KB 10.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4510_fsc.xml emd_4510_fsc.xml | 14.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_4510.png emd_4510.png | 86 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4510 http://ftp.pdbj.org/pub/emdb/structures/EMD-4510 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4510 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4510 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4510.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4510.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Listeria Stressosome
| Entire | Name: Listeria Stressosome |
|---|---|
| Components |
|
-Supramolecule #1: Listeria Stressosome
| Supramolecule | Name: Listeria Stressosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: The subcomponents are, RsbR, RsbR and he RsbT. |
|---|---|
| Source (natural) | Organism:  Listeria monocytogenes (bacteria) / Strain: EGD-e Listeria monocytogenes (bacteria) / Strain: EGD-e |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 1.8 MDa |
-Macromolecule #1: RsbR
| Macromolecule | Name: RsbR / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Source (natural) | Organism:  Listeria monocytogenes EGD-E (bacteria) Listeria monocytogenes EGD-E (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MYKDFANFIR TNKADLLNDW MNEMEKQSDQ LINDIAKEAM YEETSKEFVD LIVSNVTENG SKFNEKLDDF AEKVVHLGW PIHFVTTGLR VFGLLVYTAM RDEDLFLKRE EKPEDDAYYR FETWLSSMYN KVVTAYADTW E KTVSIQKS ALQELSAPLL PIFEKISVMP ...String: MYKDFANFIR TNKADLLNDW MNEMEKQSDQ LINDIAKEAM YEETSKEFVD LIVSNVTENG SKFNEKLDDF AEKVVHLGW PIHFVTTGLR VFGLLVYTAM RDEDLFLKRE EKPEDDAYYR FETWLSSMYN KVVTAYADTW E KTVSIQKS ALQELSAPLL PIFEKISVMP LIGTIDTERA KLIIENLLIG VVKNRSEVVL IDITGVPVVD TM VAHHIIQ ASEAVRLVGC QAMLVGIRPE IAQTIVNLGI ELDQIITTNT MKKGMERALA LTNREIVEKE G |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.02 mg/mL |
|---|---|
| Buffer | pH: 8.5 |
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 10.0 sec. / Average electron dose: 2.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)