[English] 日本語
 Yorodumi
Yorodumi- EMDB-43322: Cryo-EM structure of SARS-CoV-2 XBB.1.5 spike protein in complex ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SARS-CoV-2 XBB.1.5 spike protein in complex with mouse ACE2 (conformation 1) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationMetabolism of Angiotensinogen to Angiotensins / ceramide biosynthetic process / positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin ...Metabolism of Angiotensinogen to Angiotensins / ceramide biosynthetic process / positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction / peptidyl-dipeptidase activity / transporter activator activity / carboxypeptidase activity / angiotensin maturation / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / metallopeptidase activity / virus receptor activity / regulation of inflammatory response / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / endopeptidase activity / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / cilium / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / cell surface / negative regulation of transcription by RNA polymerase II / : / extracellular region / membrane / metal ion binding / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   | |||||||||
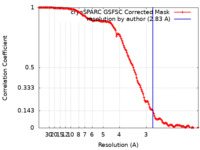
| Method | single particle reconstruction / cryo EM / Resolution: 2.83 Å | |||||||||
 Authors Authors | Zhu X / Mannar D / Saville J / Poloni C / Bezeruk A / Tidey K / Ahmed S / Tuttle K / Vahdatihassani F / Cholak S ...Zhu X / Mannar D / Saville J / Poloni C / Bezeruk A / Tidey K / Ahmed S / Tuttle K / Vahdatihassani F / Cholak S / Cook L / Steiner TS / Subramaniam S | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Altered receptor binding, antibody evasion and retention of T cell recognition by the SARS-CoV-2 XBB.1.5 spike protein. Authors: Dhiraj Mannar / James W Saville / Chad Poloni / Xing Zhu / Alison Bezeruk / Keith Tidey / Sana Ahmed / Katharine S Tuttle / Faezeh Vahdatihassani / Spencer Cholak / Laura Cook / Theodore S ...Authors: Dhiraj Mannar / James W Saville / Chad Poloni / Xing Zhu / Alison Bezeruk / Keith Tidey / Sana Ahmed / Katharine S Tuttle / Faezeh Vahdatihassani / Spencer Cholak / Laura Cook / Theodore S Steiner / Sriram Subramaniam /   Abstract: The XBB.1.5 variant of SARS-CoV-2 has rapidly achieved global dominance and exhibits a high growth advantage over previous variants. Preliminary reports suggest that the success of XBB.1.5 stems from ...The XBB.1.5 variant of SARS-CoV-2 has rapidly achieved global dominance and exhibits a high growth advantage over previous variants. Preliminary reports suggest that the success of XBB.1.5 stems from mutations within its spike glycoprotein, causing immune evasion and enhanced receptor binding. We present receptor binding studies that demonstrate retention of binding contacts with the human ACE2 receptor and a striking decrease in binding to mouse ACE2 due to the revertant R493Q mutation. Despite extensive evasion of antibody binding, we highlight a region on the XBB.1.5 spike protein receptor binding domain (RBD) that is recognized by serum antibodies from a donor with hybrid immunity, collected prior to the emergence of the XBB.1.5 variant. T cell assays reveal high frequencies of XBB.1.5 spike-specific CD4 and CD8 T cells amongst donors with hybrid immunity, with the CD4 T cells skewed towards a Th1 cell phenotype and having attenuated effector cytokine secretion as compared to ancestral spike protein-specific cells. Thus, while the XBB.1.5 variant has retained efficient human receptor binding and gained antigenic alterations, it remains susceptible to recognition by T cells induced via vaccination and previous infection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43322.map.gz emd_43322.map.gz | 123.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43322-v30.xml emd-43322-v30.xml emd-43322.xml emd-43322.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43322_fsc.xml emd_43322_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_43322.png emd_43322.png | 22.9 KB | ||
| Filedesc metadata |  emd-43322.cif.gz emd-43322.cif.gz | 7.6 KB | ||
| Others |  emd_43322_half_map_1.map.gz emd_43322_half_map_1.map.gz emd_43322_half_map_2.map.gz emd_43322_half_map_2.map.gz | 226.6 MB 226.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43322 http://ftp.pdbj.org/pub/emdb/structures/EMD-43322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43322 | HTTPS FTP |
-Related structure data
| Related structure data |  8vkmMC  8vkkC  8vklC  8vknC  8vkoC  8vkpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43322.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43322.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_43322_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_43322_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 XBB.1.5 spike protein trimer bound to mouse ACE2
| Entire | Name: SARS-CoV-2 XBB.1.5 spike protein trimer bound to mouse ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 XBB.1.5 spike protein trimer bound to mouse ACE2
| Supramolecule | Name: SARS-CoV-2 XBB.1.5 spike protein trimer bound to mouse ACE2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.226516 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPA LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYQKNNKSWM ESEFRVYSSA N NCTFEYVS ...String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPA LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYQKNNKSWM ESEFRVYSSA N NCTFEYVS QPFLMDLEGK EGNFKNLREF VFKNIDGYFK IYSKHTPINL ERDLPQGFSA LEPLVDLPIG INITRFQTLL AL HRSYLTP VDSSSGWTAG AAAYYVGYLQ PRTFLLKYNE NGTITDAVDC ALDPLSETKC TLKSFTVEKG IYQTSNFRVQ PTE SIVRFP NITNLCPFHE VFNATTFASV YAWNRKRISN CVADYSVIYN FAPFFAFKCY GVSPTKLNDL CFTNVYADSF VIRG NEVSQ IAPGQTGNIA DYNYKLPDDF TGCVIAWNSN KLDSKPSGNY NYLYRLFRKS KLKPFERDIS TEIYQAGNKP CNGVA GPNC YSPLQSYGFR PTYGVGHQPY RVVVLSFELL HAPATVCGPK KSTNLVKNKC VNFNFNGLTG TGVLTESNKK FLPFQQ FGR DIADTTDAVR DPQTLEILDI TPCSFGGVSV ITPGTNTSNQ VAVLYQGVNC TEVPVAIHAD QLTPTWRVYS TGSNVFQ TR AGCLIGAEYV NNSYECDIPI GAGICASYQT QTKSHGSASS VASQSIIAYT MSLGAENSVA YSNNSIAIPT NFTISVTT E ILPVSMTKTS VDCTMYICGD STECSNLLLQ YGSFCTQLKR ALTGIAVEQD KNTQEVFAQV KQIYKTPPIK YFGGFNFSQ ILPDPSKPSK RSPIEDLLFN KVTLADAGFI KQYGDCLGDI AARDLICAQK FNGLTVLPPL LTDEMIAQYT SALLAGTITS GWTFGAGPA LQIPFPMQMA YRFNGIGVTQ NVLYENQKLI ANQFNSAIGK IQDSLSSTPS ALGKLQDVVN HNAQALNTLV K QLSSKFGA ISSVLNDILS RLDPPEAEVQ IDRLITGRLQ SLQTYVTQQL IRAAEIRASA NLAATKMSEC VLGQSKRVDF CG KGYHLMS FPQSAPHGVV FLHVTYVPAQ EKNFTTAPAI CHDGKAHFPR EGVFVSNGTH WFVTQRNFYE PQIITTDNTF VSG NCDVVI GIVNNTVYDP LQPELDSFKE ELDKYFKNHT SPDVDLGDIS GINASVVNIQ KEIDRLNEVA KNLNESLIDL QELG KYEQG SGYIPEAPRD GQAYVRKDGE WVLLSTFLGR SLEVLFQGPG HHHHHHHHSA WSHPQFEKGG GSGGGGSGGS AWSHP QFEK UniProtKB: Spike glycoprotein |
-Macromolecule #2: Angiotensin-converting enzyme 2
| Macromolecule | Name: Angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: angiotensin-converting enzyme 2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 71.795523 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSSSSWLLLS LVAVTTAQSL TEENAKTFLN NFNQEAEDLS YQSSLASWNY NTNITEENAQ KMSEAAAKWS AFYEEQSKTA QSFSLQEIQ TPIIKRQLQA LQQSGSSALS ADKNKQLNTI LNTMSTIYST GKVCNPKNPQ ECLLLEPGLD EIMATSTDYN S RLWAWEGW ...String: MSSSSWLLLS LVAVTTAQSL TEENAKTFLN NFNQEAEDLS YQSSLASWNY NTNITEENAQ KMSEAAAKWS AFYEEQSKTA QSFSLQEIQ TPIIKRQLQA LQQSGSSALS ADKNKQLNTI LNTMSTIYST GKVCNPKNPQ ECLLLEPGLD EIMATSTDYN S RLWAWEGW RAEVGKQLRP LYEEYVVLKN EMARANNYND YGDYWRGDYE AEGADGYNYN RNQLIEDVER TFAEIKPLYE HL HAYVRRK LMDTYPSYIS PTGCLPAHLL GDMWGRFWTN LYPLTVPFAQ KPNIDVTDAM MNQGWDAERI FQEAEKFFVS VGL PHMTQG FWANSMLTEP ADGRKVVCHP TAWDLGHGDF RIKMCTKVTM DNFLTAHHEM GHIQYDMAYA RQPFLLRNGA NEGF HEAVG EIMSLSAATP KHLKSIGLLP SDFQEDSETE INFLLKQALT IVGTLPFTYM LEKWRWMVFR GEIPKEQWMK KWWEM KREI VGVVEPLPHD ETYCDPASLF HVSNDYSFIR YYTRTIYQFQ FQEALCQAAK YNGSLHKCDI SNSTEAGQKL LKMLSL GNS EPWTKALENV VGARNMDVKP LLNYFQPLFD WLKEQNRNSF VGWNTEWSPY ADHHHHHH UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)