[English] 日本語
 Yorodumi
Yorodumi- EMDB-43195: Cryo-EM structure of FoxA1 in complex with ALBN1 nucleosome (class 2) -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of FoxA1 in complex with ALBN1 nucleosome (class 2) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nucleosome / pioneer transcription factors / DNA binding proteins / transcription / chromatin / NUCLEAR PROTEIN / NUCLEAR PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationalveolar secondary septum development / respiratory basal cell differentiation / positive regulation of dopaminergic neuron differentiation / mesenchymal-epithelial cell signaling involved in prostate gland development / anatomical structure formation involved in morphogenesis / prostate gland stromal morphogenesis / positive regulation of cell-cell adhesion mediated by cadherin / epithelial cell maturation involved in prostate gland development / neuron fate specification / lung epithelial cell differentiation ...alveolar secondary septum development / respiratory basal cell differentiation / positive regulation of dopaminergic neuron differentiation / mesenchymal-epithelial cell signaling involved in prostate gland development / anatomical structure formation involved in morphogenesis / prostate gland stromal morphogenesis / positive regulation of cell-cell adhesion mediated by cadherin / epithelial cell maturation involved in prostate gland development / neuron fate specification / lung epithelial cell differentiation / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / prostate gland epithelium morphogenesis / dopaminergic neuron differentiation / Formation of axial mesoderm / positive regulation of smoothened signaling pathway / hormone metabolic process / negative regulation of epithelial to mesenchymal transition / smoothened signaling pathway / dorsal/ventral neural tube patterning / positive regulation of intracellular estrogen receptor signaling pathway / epithelial tube branching involved in lung morphogenesis / microvillus / : / anatomical structure morphogenesis / negative regulation of tumor necrosis factor-mediated signaling pathway / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Notch signaling pathway / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / positive regulation of mitotic cell cycle / Deposition of new CENPA-containing nucleosomes at the centromere / epigenetic regulation of gene expression / telomere organization / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Interleukin-7 signaling / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / innate immune response in mucosa / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / lipopolysaccharide binding / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / NoRC negatively regulates rRNA expression / positive regulation of miRNA transcription / B-WICH complex positively regulates rRNA expression / PKMTs methylate histone lysines / DNA Damage/Telomere Stress Induced Senescence / Pre-NOTCH Transcription and Translation / Meiotic recombination / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Metalloprotease DUBs / fibrillar center / Transcriptional regulation of granulopoiesis / RMTs methylate histone arginines / HCMV Early Events / sequence-specific double-stranded DNA binding / structural constituent of chromatin / UCH proteinases / response to estradiol / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / heterochromatin formation / nucleosome assembly / glucose homeostasis / E3 ubiquitin ligases ubiquitinate target proteins / antibacterial humoral response / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / HATs acetylate histones / Factors involved in megakaryocyte development and platelet production / RUNX1 regulates transcription of genes involved in differentiation of HSCs / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / chromatin organization / Processing of DNA double-strand break ends Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
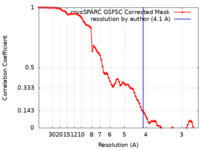
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Zhou BR / Bai Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2024 Journal: Mol Cell / Year: 2024Title: Structural insights into the cooperative nucleosome recognition and chromatin opening by FOXA1 and GATA4. Authors: Bing-Rui Zhou / Hanqiao Feng / Furong Huang / Iris Zhu / Stephanie Portillo-Ledesma / Dan Shi / Kenneth S Zaret / Tamar Schlick / David Landsman / Qianben Wang / Yawen Bai /   Abstract: Mouse FOXA1 and GATA4 are prototypes of pioneer factors, initiating liver cell development by binding to the N1 nucleosome in the enhancer of the ALB1 gene. Using cryoelectron microscopy (cryo-EM), ...Mouse FOXA1 and GATA4 are prototypes of pioneer factors, initiating liver cell development by binding to the N1 nucleosome in the enhancer of the ALB1 gene. Using cryoelectron microscopy (cryo-EM), we determined the structures of the free N1 nucleosome and its complexes with FOXA1 and GATA4, both individually and in combination. We found that the DNA-binding domains of FOXA1 and GATA4 mainly recognize the linker DNA and an internal site in the nucleosome, respectively, whereas their intrinsically disordered regions interact with the acidic patch on histone H2A-H2B. FOXA1 efficiently enhances GATA4 binding by repositioning the N1 nucleosome. In vivo DNA editing and bioinformatics analyses suggest that the co-binding mode of FOXA1 and GATA4 plays important roles in regulating genes involved in liver cell functions. Our results reveal the mechanism whereby FOXA1 and GATA4 cooperatively bind to the nucleosome through nucleosome repositioning, opening chromatin by bending linker DNA and obstructing nucleosome packing. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43195.map.gz emd_43195.map.gz | 32.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43195-v30.xml emd-43195-v30.xml emd-43195.xml emd-43195.xml | 25.6 KB 25.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43195_fsc.xml emd_43195_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_43195.png emd_43195.png | 99.8 KB | ||
| Masks |  emd_43195_msk_1.map emd_43195_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-43195.cif.gz emd-43195.cif.gz | 7.6 KB | ||
| Others |  emd_43195_half_map_1.map.gz emd_43195_half_map_1.map.gz emd_43195_half_map_2.map.gz emd_43195_half_map_2.map.gz | 59.5 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43195 http://ftp.pdbj.org/pub/emdb/structures/EMD-43195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43195 | HTTPS FTP |
-Related structure data
| Related structure data |  8vfzMC  8vfxC  8vfyC  8vg0C  8vg1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43195.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43195.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||
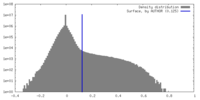
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_43195_msk_1.map emd_43195_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
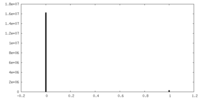
| Density Histograms |
-Half map: #2
| File | emd_43195_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
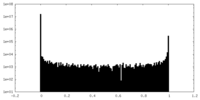
| Density Histograms |
-Half map: #1
| File | emd_43195_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
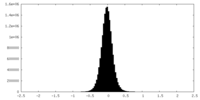
| Density Histograms |
- Sample components
Sample components
-Entire : FoxA1 in complx with 186bp ALBN1 nucleosome
| Entire | Name: FoxA1 in complx with 186bp ALBN1 nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: FoxA1 in complx with 186bp ALBN1 nucleosome
| Supramolecule | Name: FoxA1 in complx with 186bp ALBN1 nucleosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Hepatocyte nuclear factor 3-alpha
| Macromolecule | Name: Hepatocyte nuclear factor 3-alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.022562 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLGTVKMEGH ETSDWNSYYA DTQEAYSSVP VSNMNSGLGS MNSMNTYMTM NTMTTSGNMT PASFNMSYAN PGLGAGLSPG AVAGMPGGS AGAMNSMTAA GVTAMGTALS PSGMGAMGAQ QAASMNGLGP YAAAMNPCMS PMAYAPSNLG RSRAGGGGDA K TFKRSYPH ...String: MLGTVKMEGH ETSDWNSYYA DTQEAYSSVP VSNMNSGLGS MNSMNTYMTM NTMTTSGNMT PASFNMSYAN PGLGAGLSPG AVAGMPGGS AGAMNSMTAA GVTAMGTALS PSGMGAMGAQ QAASMNGLGP YAAAMNPCMS PMAYAPSNLG RSRAGGGGDA K TFKRSYPH AKPPYSYISL ITMAIQQAPS KMLTLSEIYQ WIMDLFPYYR QNQQRWQNSI RHSLSFNDCF VKVARSPDKP GK GSYWTLH PDSGNMFENG CYLRRQKRFK CEKQPGAGGG GGSGSGGSGA KGGPESRKDP SGASNPSADS PLHRGVHGKT GQL EGAPAP GPAASPQTLD HSGATATGGA SELKTPASST APPISSGPGA LASVPASHPA HGLAPHESQL HLKGDPHYSF NHPF SINNL MSSSEQQHKL DFKAYEQALQ YSPYGSTLPA SLPLGSASVT TRSPIEPSAL EPAYYQGVYS RPVLNTSHHH HHH UniProtKB: Hepatocyte nuclear factor 3-alpha |
-Macromolecule #4: Histone H2A type 1-B/E
| Macromolecule | Name: Histone H2A type 1-B/E / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.165551 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKQGGK ARAKAKTRSS RAGLQFPVGR VHRLLRKGNY SERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A type 1-B/E |
-Macromolecule #5: Histone H3.1
| Macromolecule | Name: Histone H3.1 / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.437167 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEACEA YLVGLFEDTN LCAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.1 |
-Macromolecule #6: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.394426 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-Macromolecule #7: Histone H2B type 1-J
| Macromolecule | Name: Histone H2B type 1-J / type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.935239 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSI YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSAK UniProtKB: Histone H2B type 1-J |
-Macromolecule #2: DNA (171-MER)
| Macromolecule | Name: DNA (171-MER) / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 57.399637 KDa |
| Sequence | String: (DA)(DT)(DC)(DC)(DG)(DA)(DG)(DA)(DT)(DG) (DG)(DT)(DA)(DC)(DT)(DT)(DT)(DG)(DT)(DG) (DT)(DC)(DT)(DC)(DC)(DT)(DG)(DC)(DT) (DC)(DT)(DG)(DT)(DC)(DA)(DG)(DC)(DA)(DG) (DG) (DG)(DC)(DA)(DC)(DT)(DG) ...String: (DA)(DT)(DC)(DC)(DG)(DA)(DG)(DA)(DT)(DG) (DG)(DT)(DA)(DC)(DT)(DT)(DT)(DG)(DT)(DG) (DT)(DC)(DT)(DC)(DC)(DT)(DG)(DC)(DT) (DC)(DT)(DG)(DT)(DC)(DA)(DG)(DC)(DA)(DG) (DG) (DG)(DC)(DA)(DC)(DT)(DG)(DT)(DA) (DC)(DT)(DT)(DG)(DC)(DT)(DG)(DA)(DT)(DA) (DC)(DC) (DA)(DG)(DG)(DG)(DA)(DA)(DT) (DG)(DT)(DT)(DT)(DG)(DT)(DT)(DC)(DT)(DT) (DA)(DA)(DA) (DT)(DA)(DC)(DC)(DA)(DT) (DC)(DA)(DT)(DT)(DC)(DC)(DG)(DG)(DA)(DC) (DG)(DT)(DG)(DT) (DT)(DT)(DG)(DC)(DC) (DT)(DT)(DG)(DG)(DC)(DC)(DA)(DG)(DT)(DT) (DT)(DT)(DC)(DC)(DA) (DT)(DG)(DT)(DA) (DC)(DA)(DT)(DG)(DC)(DA)(DG)(DA)(DA)(DA) (DG)(DA)(DA)(DG)(DT)(DT) (DT)(DG)(DG) (DA)(DC)(DT)(DG)(DA)(DT)(DC)(DA)(DA)(DT) (DA)(DC)(DA)(DG)(DT)(DC)(DC) (DT)(DC) (DT)(DG)(DC)(DC)(DT)(DT)(DT)(DA)(DA)(DA) (DG)(DC)(DA)(DA)(DT)(DA)(DG)(DG) (DA) (DA)(DA)(DG)(DA)(DT) |
-Macromolecule #3: DNA (171-MER)
| Macromolecule | Name: DNA (171-MER) / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 57.42777 KDa |
| Sequence | String: (DA)(DT)(DC)(DT)(DT)(DT)(DC)(DC)(DT)(DA) (DT)(DT)(DG)(DC)(DT)(DT)(DT)(DA)(DA)(DA) (DG)(DG)(DC)(DA)(DG)(DA)(DG)(DG)(DA) (DC)(DT)(DG)(DT)(DA)(DT)(DT)(DG)(DA)(DT) (DC) (DA)(DG)(DT)(DC)(DC)(DA) ...String: (DA)(DT)(DC)(DT)(DT)(DT)(DC)(DC)(DT)(DA) (DT)(DT)(DG)(DC)(DT)(DT)(DT)(DA)(DA)(DA) (DG)(DG)(DC)(DA)(DG)(DA)(DG)(DG)(DA) (DC)(DT)(DG)(DT)(DA)(DT)(DT)(DG)(DA)(DT) (DC) (DA)(DG)(DT)(DC)(DC)(DA)(DA)(DA) (DC)(DT)(DT)(DC)(DT)(DT)(DT)(DC)(DT)(DG) (DC)(DA) (DT)(DG)(DT)(DA)(DC)(DA)(DT) (DG)(DG)(DA)(DA)(DA)(DA)(DC)(DT)(DG)(DG) (DC)(DC)(DA) (DA)(DG)(DG)(DC)(DA)(DA) (DA)(DC)(DA)(DC)(DG)(DT)(DC)(DC)(DG)(DG) (DA)(DA)(DT)(DG) (DA)(DT)(DG)(DG)(DT) (DA)(DT)(DT)(DT)(DA)(DA)(DG)(DA)(DA)(DC) (DA)(DA)(DA)(DC)(DA) (DT)(DT)(DC)(DC) (DC)(DT)(DG)(DG)(DT)(DA)(DT)(DC)(DA)(DG) (DC)(DA)(DA)(DG)(DT)(DA) (DC)(DA)(DG) (DT)(DG)(DC)(DC)(DC)(DT)(DG)(DC)(DT)(DG) (DA)(DC)(DA)(DG)(DA)(DG)(DC) (DA)(DG) (DG)(DA)(DG)(DA)(DC)(DA)(DC)(DA)(DA)(DA) (DG)(DT)(DA)(DC)(DC)(DA)(DT)(DC) (DT) (DC)(DG)(DG)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)