[English] 日本語
 Yorodumi
Yorodumi- EMDB-42515: Intracellular Ebola nucleocapsid-like structure obtained from cel... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Intracellular Ebola nucleocapsid-like structure obtained from cells expressing NP, VP24 and VP35 at 17.6 angstrom | |||||||||
 Map data Map data | intracellular Ebola virus nucelocapsid from cells expressing NP, VP24 and VP35 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | VIRAL PROTEIN / nucleoprotein / nucleocapsid / Ebola virus / EBOV / filovirus / subtomogram averaging / cryo-ET / FIB / intracellular / in situ | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 17.57 Å | |||||||||
 Authors Authors | Watanabe R / Zyla D / Saphire EO | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: Intracellular Ebola virus nucleocapsid assembly revealed by in situ cryo-electron tomography. Authors: Reika Watanabe / Dawid Zyla / Diptiben Parekh / Connor Hong / Ying Jones / Sharon L Schendel / William Wan / Guillaume Castillon / Erica Ollmann Saphire /  Abstract: Filoviruses, including the Ebola and Marburg viruses, cause hemorrhagic fevers with up to 90% lethality. The viral nucleocapsid is assembled by polymerization of the nucleoprotein (NP) along the ...Filoviruses, including the Ebola and Marburg viruses, cause hemorrhagic fevers with up to 90% lethality. The viral nucleocapsid is assembled by polymerization of the nucleoprotein (NP) along the viral genome, together with the viral proteins VP24 and VP35. We employed cryo-electron tomography of cells transfected with viral proteins and infected with model Ebola virus to illuminate assembly intermediates, as well as a 9 Å map of the complete intracellular assembly. This structure reveals a previously unresolved third and outer layer of NP complexed with VP35. The intrinsically disordered region, together with the C-terminal domain of this outer layer of NP, provides the constant width between intracellular nucleocapsid bundles and likely functions as a flexible tether to the viral matrix protein in the virion. A comparison of intracellular nucleocapsids with prior in-virion nucleocapsid structures reveals that the nucleocapsid further condenses vertically in the virion. The interfaces responsible for nucleocapsid assembly are highly conserved and offer targets for broadly effective antivirals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42515.map.gz emd_42515.map.gz | 580.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42515-v30.xml emd-42515-v30.xml emd-42515.xml emd-42515.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_42515.png emd_42515.png | 94.7 KB | ||
| Masks |  emd_42515_msk_1.map emd_42515_msk_1.map | 1.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-42515.cif.gz emd-42515.cif.gz | 4.4 KB | ||
| Others |  emd_42515_half_map_1.map.gz emd_42515_half_map_1.map.gz emd_42515_half_map_2.map.gz emd_42515_half_map_2.map.gz | 1 MB 1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42515 http://ftp.pdbj.org/pub/emdb/structures/EMD-42515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42515 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42515.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42515.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | intracellular Ebola virus nucelocapsid from cells expressing NP, VP24 and VP35 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.37 Å | ||||||||||||||||||||||||||||||||||||
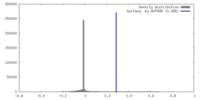
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_42515_msk_1.map emd_42515_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
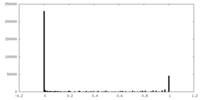
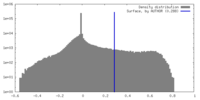
| Density Histograms |
-Half map: half map1
| File | emd_42515_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
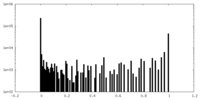
| Density Histograms |
-Half map: half map2
| File | emd_42515_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
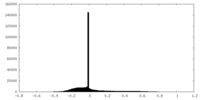
| Density Histograms |
- Sample components
Sample components
-Entire : Ebola virus - Mayinga, Zaire, 1976
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Ebola virus - Mayinga, Zaire, 1976
| Supramolecule | Name: Ebola virus - Mayinga, Zaire, 1976 / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 128952 / Sci species name: Ebola virus - Mayinga, Zaire, 1976 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE-PROPANE |
| Details | FIB-milled-plunge-frozen cell expressing EBOV NP, VP24 and VP35 |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 3.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.5 µm / Nominal defocus min: 4.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 17.57 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number subtomograms used: 8371 |
|---|---|
| Extraction | Number tomograms: 10 / Number images used: 39000 |
| Final 3D classification | Number classes: 3 / Avg.num./class: 13000 / Software - Name: RELION (ver. 3.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
 Movie
Movie Controller
Controller


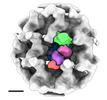



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)