+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Firmicutes Rubisco | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Carboxylase / Oxygenase / LYASE | |||||||||
| Biological species |  Bacillota (low GC Gram+) Bacillota (low GC Gram+) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.21 Å | |||||||||
 Authors Authors | Kaeser BP / Liu AK / Shih PM | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Curr Biol / Year: 2023 Journal: Curr Biol / Year: 2023Title: Deep-branching evolutionary intermediates reveal structural origins of form I rubisco. Authors: Albert K Liu / Benjamin Kaeser / LinXing Chen / Jacob West-Roberts / Leah J Taylor-Kearney / Adi Lavy / Damian Günzing / Wen-Jun Li / Michal Hammel / Eva Nogales / Jillian F Banfield / Patrick M Shih /    Abstract: The enzyme rubisco (ribulose-1,5-bisphosphate carboxylase/oxygenase) catalyzes the majority of biological carbon fixation on Earth. Although the vast majority of rubiscos across the tree of life ...The enzyme rubisco (ribulose-1,5-bisphosphate carboxylase/oxygenase) catalyzes the majority of biological carbon fixation on Earth. Although the vast majority of rubiscos across the tree of life assemble as homo-oligomers, the globally predominant form I enzyme-found in plants, algae, and cyanobacteria-forms a unique hetero-oligomeric complex. The recent discovery of a homo-oligomeric sister group to form I rubisco (named form I') has filled a key gap in our understanding of the enigmatic origins of the form I clade. However, to elucidate the series of molecular events leading to the evolution of form I rubisco, we must examine more distantly related sibling clades to contextualize the molecular features distinguishing form I and form I' rubiscos. Here, we present a comparative structural study retracing the evolutionary history of rubisco that reveals a complex structural trajectory leading to the ultimate hetero-oligomerization of the form I clade. We structurally characterize the oligomeric states of deep-branching form Iα and I'' rubiscos recently discovered from metagenomes, which represent key evolutionary intermediates preceding the form I clade. We further solve the structure of form I'' rubisco, revealing the molecular determinants that likely primed the enzyme core for the transition from a homo-oligomer to a hetero-oligomer. Our findings yield new insight into the evolutionary trajectory underpinning the adoption and entrenchment of the prevalent assembly of form I rubisco, providing additional context when viewing the enzyme family through the broader lens of protein evolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41946.map.gz emd_41946.map.gz | 97.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41946-v30.xml emd-41946-v30.xml emd-41946.xml emd-41946.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
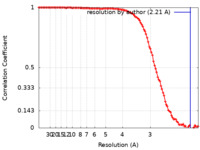
| FSC (resolution estimation) |  emd_41946_fsc.xml emd_41946_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_41946.png emd_41946.png | 39.6 KB | ||
| Filedesc metadata |  emd-41946.cif.gz emd-41946.cif.gz | 6.4 KB | ||
| Others |  emd_41946_half_map_1.map.gz emd_41946_half_map_1.map.gz emd_41946_half_map_2.map.gz emd_41946_half_map_2.map.gz | 95.3 MB 95.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41946 http://ftp.pdbj.org/pub/emdb/structures/EMD-41946 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41946 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41946 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41946.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41946.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41946_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
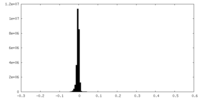
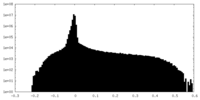
| Density Histograms |
-Half map: #1
| File | emd_41946_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
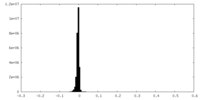
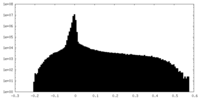
| Density Histograms |
- Sample components
Sample components
-Entire : RUBISCO octamer
| Entire | Name: RUBISCO octamer |
|---|---|
| Components |
|
-Supramolecule #1: RUBISCO octamer
| Supramolecule | Name: RUBISCO octamer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Bacillota (low GC Gram+) Bacillota (low GC Gram+) |
-Macromolecule #1: Rubisco
| Macromolecule | Name: Rubisco / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bacillota (low GC Gram+) Bacillota (low GC Gram+) |
| Molecular weight | Theoretical: 50.978855 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARQQFVAGV QPYRKTYYEP GYEPKETDLL CAFRIEPSPG IPLEEAAAAV AAESSTGTWT EVWSQEMTDL HRYKGRCYAI DGNTAYIAY PLDLFEEGSI VNVMSSIVGN VFGFKAVRAL RLLDMRIPTA YLKTFPGPPT GIAQERDRLK VYHRPLLGGT I KPKLGLGP ...String: MARQQFVAGV QPYRKTYYEP GYEPKETDLL CAFRIEPSPG IPLEEAAAAV AAESSTGTWT EVWSQEMTDL HRYKGRCYAI DGNTAYIAY PLDLFEEGSI VNVMSSIVGN VFGFKAVRAL RLLDMRIPTA YLKTFPGPPT GIAQERDRLK VYHRPLLGGT I KPKLGLGP KEFARVVYEC LVGGLDTT(KCX)D DENLNSQPFC RWRDRYLYVM DAVHRAEEET GEAKGHWLNV TAGDTEEM L RRAEFAKEVG SRYIMVDFLT AGFSAYATLR RRAEELGLMI HCHRAMHAVF TRPKDHGIHF RVVAKWLRMA GGDHVHTGT VVGKLEGARE EVRGIADLLR EEFVPANPQR GLLFDQPWAG LKPLFPVASG GIHVWHVPDL VSIYGNDAFF LFGGGTHGHP RGSRAGARA NRAAVEAVAA AYREGRDILA EGRQILQDAA RTCPELREAM ELWEGVTFGE E |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: 2-CARBOXYARABINITOL-1,5-DIPHOSPHATE
| Macromolecule | Name: 2-CARBOXYARABINITOL-1,5-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 8 / Formula: CAP |
|---|---|
| Molecular weight | Theoretical: 356.115 Da |
| Chemical component information |  ChemComp-CAP: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 981 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)