[English] 日本語
 Yorodumi
Yorodumi- EMDB-41740: Cryo-EM reconstruction of bovine concentrative nucleoside transpo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of bovine concentrative nucleoside transporter 3 in complex with Molnupiravir, condition 1, consensus reconstruction | |||||||||
 Map data Map data | full map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | membrane protein / transporter / nucleoside / TRANSPORT PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Wright NJ / Lee S-Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2024 Journal: Nat Chem Biol / Year: 2024Title: Antiviral drug recognition and elevator-type transport motions of CNT3. Authors: Nicholas J Wright / Feng Zhang / Yang Suo / Lingyang Kong / Ying Yin / Justin G Fedor / Kedar Sharma / Mario J Borgnia / Wonpil Im / Seok-Yong Lee /  Abstract: Nucleoside analogs have broad clinical utility as antiviral drugs. Key to their systemic distribution and cellular entry are human nucleoside transporters. Here, we establish that the human ...Nucleoside analogs have broad clinical utility as antiviral drugs. Key to their systemic distribution and cellular entry are human nucleoside transporters. Here, we establish that the human concentrative nucleoside transporter 3 (CNT3) interacts with antiviral drugs used in the treatment of coronavirus infections. We report high-resolution single-particle cryo-electron microscopy structures of bovine CNT3 complexed with antiviral nucleosides N-hydroxycytidine, PSI-6206, GS-441524 and ribavirin, all in inward-facing states. Notably, we found that the orally bioavailable antiviral molnupiravir arrests CNT3 in four distinct conformations, allowing us to capture cryo-electron microscopy structures of drug-loaded outward-facing and drug-loaded intermediate states. Our studies uncover the conformational trajectory of CNT3 during membrane transport of a nucleoside analog antiviral drug, yield new insights into the role of interactions between the transport and the scaffold domains in elevator-like domain movements during drug translocation, and provide insights into the design of nucleoside analog antiviral prodrugs with improved oral bioavailability. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41740.map.gz emd_41740.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41740-v30.xml emd-41740-v30.xml emd-41740.xml emd-41740.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41740.png emd_41740.png | 55.6 KB | ||
| Filedesc metadata |  emd-41740.cif.gz emd-41740.cif.gz | 5.8 KB | ||
| Others |  emd_41740_half_map_1.map.gz emd_41740_half_map_1.map.gz emd_41740_half_map_2.map.gz emd_41740_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41740 http://ftp.pdbj.org/pub/emdb/structures/EMD-41740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41740 | HTTPS FTP |
-Related structure data
| Related structure data |  8tz1C  8tz2C  8tz3C  8tz4C  8tz5C  8tz6C  8tz7C  8tz8C  8tz9C  8tzaC  8tzdC C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41740.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41740.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | full map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
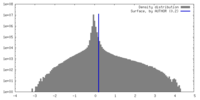
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 1
| File | emd_41740_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
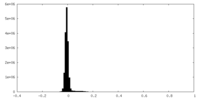
| Density Histograms |
- Sample components
Sample components
-Entire : bCNT3 trimer
| Entire | Name: bCNT3 trimer |
|---|---|
| Components |
|
-Supramolecule #1: bCNT3 trimer
| Supramolecule | Name: bCNT3 trimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 232 KDa |
-Macromolecule #1: bCNT3
| Macromolecule | Name: bCNT3 / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MDYKDDDDKL EATMAMSSKI SVELQRVAAL PAQGCSNTGF QNDEDGFENQ NPSGNDHSLR NRVVQNREHE NGKQVEEHIT IGQDSLRKDE EEEDDQETHR KGCLERMCGR MSDFCREHKT TLRYIIWGIL IAGYLALVIA ACVMNFHRAL PLFVITVVAI FFVVWDHLMA ...String: MDYKDDDDKL EATMAMSSKI SVELQRVAAL PAQGCSNTGF QNDEDGFENQ NPSGNDHSLR NRVVQNREHE NGKQVEEHIT IGQDSLRKDE EEEDDQETHR KGCLERMCGR MSDFCREHKT TLRYIIWGIL IAGYLALVIA ACVMNFHRAL PLFVITVVAI FFVVWDHLMA KYESQIARFL SPGQRLLDSH WFWLKWVIWG CLILGVILWL VFDTAKLGQQ QLVSFGGLII YTSLTFLFSK HPTKVYWRPV FWGIGLQFLL GLLILRTEPG FMAFDWLGKQ VQTFLGYSDA GASFVFGEKY TDHFFAFKVL PIVIFFSTVM SMLYYLGLMQ WIIRKVGWVM LVTMGTSPVE SVVASGNIFI GQTESPLLVR PYLPYVTKSE LHAIMTAGFS TIAGSVLGAY ISFGVSSSHL LTASVMSAPA ALAISKLFWP ETETPKINLK NAMKMESGDS RNLLEAATQG ASSSISLVAN IAVNLIAFLA LLSFMNSALS WLGNMFDYPQ LSFEVICSYV FMPFAFMMGV DWQDSFMVAK LIGYKTFFNE FVAYQQLSKL ISLRQVGGPK FVDGVQQYMS MRSEAISTYA LCGFANFGSL GIVIGGLTSM APSRKRDITA GAMRALIAGT IACFLTACIA GMLTNTPVDI NCHHILENAF NSGLVRNTTN VVSCCQGLLS SAVVKGPGEV IPTGNHSLYS LKNCCNLLNT PTLNCSWIPN VLSNS GENBANK: GENBANK: F1MGR1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 / Component:
| ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 4715 / Average exposure time: 2.4 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)