[English] 日本語
 Yorodumi
Yorodumi- EMDB-40905: Cryo-EM Structure of NINJ1 Filament at 2.75 Angstrom Resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of NINJ1 Filament at 2.75 Angstrom Resolution | |||||||||
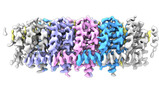
 Map data Map data | Cryo-EM Map of NINJ1 Filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NINJ1 Filament / Plasma Membrane Rupture Protein / Cholesterol Binding Protein / Lipid Binding Protein / Membrane Protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationferroptosis / membrane destabilizing activity / pyroptotic cell death / cytolysis / leukocyte chemotaxis involved in inflammatory response / positive regulation of toll-like receptor 4 signaling pathway / cell adhesion mediator activity / programmed necrotic cell death / tissue regeneration / muscle cell differentiation ...ferroptosis / membrane destabilizing activity / pyroptotic cell death / cytolysis / leukocyte chemotaxis involved in inflammatory response / positive regulation of toll-like receptor 4 signaling pathway / cell adhesion mediator activity / programmed necrotic cell death / tissue regeneration / muscle cell differentiation / cellular hyperosmotic response / heterotypic cell-cell adhesion / synaptic membrane / lipopolysaccharide binding / protein homooligomerization / positive regulation of angiogenesis / positive regulation of inflammatory response / nervous system development / angiogenesis / killing of cells of another organism / cell adhesion / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
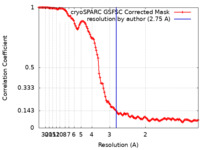
| Method | single particle reconstruction / cryo EM / Resolution: 2.75 Å | |||||||||
 Authors Authors | Sahoo B / Dai X | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2025 Journal: Cell / Year: 2025Title: How NINJ1 mediates plasma membrane rupture and why NINJ2 cannot. Authors: Bibekananda Sahoo / Zongjun Mou / Wei Liu / George Dubyak / Xinghong Dai /  Abstract: Ninjurin-1 (NINJ1) is an active executioner of plasma membrane rupture (PMR), a process previously thought to be a passive osmotic lysis event in lytic cell death. Ninjurin-2 (NINJ2) is a close ...Ninjurin-1 (NINJ1) is an active executioner of plasma membrane rupture (PMR), a process previously thought to be a passive osmotic lysis event in lytic cell death. Ninjurin-2 (NINJ2) is a close paralog of NINJ1 but cannot mediate PMR. Using cryogenic electron microscopy (cryo-EM), we show that NINJ1 and NINJ2 both assemble into linear filaments that are hydrophobic on one side but hydrophilic on the other. This structural feature and other evidence point to a PMR mechanism by which NINJ1 filaments wrap around and solubilize membrane fragments and, less frequently, form pores in the plasma membrane. In contrast to the straight NINJ1 filament, the NINJ2 filament is curved toward the intracellular space, preventing its circularization or even assembly on a relatively flat membrane to mediate PMR. Mutagenesis studies further demonstrate that the NINJ2 filament curvature is induced by strong association with lipids, particularly a cholesterol molecule, at the cytoplasmic leaflet of the lipid bilayer. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40905.map.gz emd_40905.map.gz | 168 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40905-v30.xml emd-40905-v30.xml emd-40905.xml emd-40905.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40905_fsc.xml emd_40905_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_40905.png emd_40905.png | 96.7 KB | ||
| Filedesc metadata |  emd-40905.cif.gz emd-40905.cif.gz | 5.7 KB | ||
| Others |  emd_40905_half_map_1.map.gz emd_40905_half_map_1.map.gz emd_40905_half_map_2.map.gz emd_40905_half_map_2.map.gz | 165.4 MB 165.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40905 http://ftp.pdbj.org/pub/emdb/structures/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40905 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40905 | HTTPS FTP |
-Related structure data
| Related structure data |  8szaMC  8szbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40905.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40905.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Map of NINJ1 Filament | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.66 Å | ||||||||||||||||||||||||||||||||||||
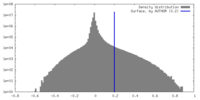
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Cryo-EM Half Map A of NINJ1 Filament
| File | emd_40905_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Half Map A of NINJ1 Filament | ||||||||||||
| Projections & Slices |
| ||||||||||||
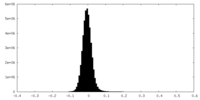
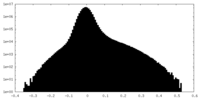
| Density Histograms |
-Half map: Cryo-EM Half Map B of NINJ1 Filament
| File | emd_40905_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM Half Map B of NINJ1 Filament | ||||||||||||
| Projections & Slices |
| ||||||||||||
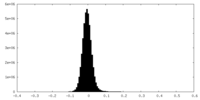
| Density Histograms |
- Sample components
Sample components
-Entire : Ninjurin-1 in complex with Cholesterol
| Entire | Name: Ninjurin-1 in complex with Cholesterol |
|---|---|
| Components |
|
-Supramolecule #1: Ninjurin-1 in complex with Cholesterol
| Supramolecule | Name: Ninjurin-1 in complex with Cholesterol / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Ninjurin-1
| Macromolecule | Name: Ninjurin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 19.276896 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDHDGDY KDHDIDYKDD DDKGSGDSGT EEYELNGGLP PGTPGSPDAS PARWGWRHGP INVNHYASKK SAAESMLDIA LLMANASQL KAVVEQGPSF AFYVPLVVLI SISLVLQIGV GVLLIFLVKY DLNNPAKHAK LDFLNNLATG LVFIIVVVNI F ITAFGVQK PLMDMAPQQ UniProtKB: Ninjurin-1 |
-Macromolecule #2: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 2 / Number of copies: 6 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)