[English] 日本語
 Yorodumi
Yorodumi- EMDB-40765: Cryo-EM structure of the wild type P. aeruginosa flagellar filament -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the wild type P. aeruginosa flagellar filament | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacterial flagellar filament / motility / Pseudomonas / STRUCTURAL PROTEIN / PROTEIN FILAMENT | |||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum / bacterial-type flagellum-dependent cell motility / structural molecule activity / extracellular region Similarity search - Function | |||||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) | |||||||||
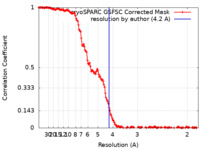
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Kreutzberger MA / Nedeljkovic M / Egelman EH / Sundberg EJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2023 Journal: PLoS Pathog / Year: 2023Title: An unbroken network of interactions connecting flagellin domains is required for motility in viscous environments. Authors: Marko Nedeljković / Mark A B Kreutzberger / Sandra Postel / Daniel Bonsor / Yingying Xing / Neil Jacob / William J Schuler / Edward H Egelman / Eric J Sundberg /   Abstract: In its simplest form, bacterial flagellar filaments are composed of flagellin proteins with just two helical inner domains, which together comprise the filament core. Although this minimal filament ...In its simplest form, bacterial flagellar filaments are composed of flagellin proteins with just two helical inner domains, which together comprise the filament core. Although this minimal filament is sufficient to provide motility in many flagellated bacteria, most bacteria produce flagella composed of flagellin proteins with one or more outer domains arranged in a variety of supramolecular architectures radiating from the inner core. Flagellin outer domains are known to be involved in adhesion, proteolysis and immune evasion but have not been thought to be required for motility. Here we show that in the Pseudomonas aeruginosa PAO1 strain, a bacterium that forms a ridged filament with a dimerization of its flagellin outer domains, motility is categorically dependent on these flagellin outer domains. Moreover, a comprehensive network of intermolecular interactions connecting the inner domains to the outer domains, the outer domains to one another, and the outer domains back to the inner domain filament core, is required for motility. This inter-domain connectivity confers PAO1 flagella with increased stability, essential for its motility in viscous environments. Additionally, we find that such ridged flagellar filaments are not unique to Pseudomonas but are, instead, present throughout diverse bacterial phyla. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40765.map.gz emd_40765.map.gz | 38.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40765-v30.xml emd-40765-v30.xml emd-40765.xml emd-40765.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40765_fsc.xml emd_40765_fsc.xml | 14.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_40765.png emd_40765.png | 49.5 KB | ||
| Masks |  emd_40765_msk_1.map emd_40765_msk_1.map | 307.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40765.cif.gz emd-40765.cif.gz | 5.2 KB | ||
| Others |  emd_40765_half_map_1.map.gz emd_40765_half_map_1.map.gz emd_40765_half_map_2.map.gz emd_40765_half_map_2.map.gz | 285 MB 285 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40765 http://ftp.pdbj.org/pub/emdb/structures/EMD-40765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40765 | HTTPS FTP |
-Validation report
| Summary document |  emd_40765_validation.pdf.gz emd_40765_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40765_full_validation.pdf.gz emd_40765_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_40765_validation.xml.gz emd_40765_validation.xml.gz | 23 KB | Display | |
| Data in CIF |  emd_40765_validation.cif.gz emd_40765_validation.cif.gz | 29.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40765 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40765 | HTTPS FTP |
-Related structure data
| Related structure data |  8sugMC  8ermC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40765.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40765.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40765_msk_1.map emd_40765_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
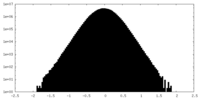
| Density Histograms |
-Half map: #2
| File | emd_40765_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_40765_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bacterial flagellar filament
| Entire | Name: Bacterial flagellar filament |
|---|---|
| Components |
|
-Supramolecule #1: Bacterial flagellar filament
| Supramolecule | Name: Bacterial flagellar filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
-Macromolecule #1: B-type flagellin
| Macromolecule | Name: B-type flagellin / type: protein_or_peptide / ID: 1 / Number of copies: 33 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 48.872355 KDa |
| Sequence | String: VNTNIASLNT QRNLNASSND LNTSLQRLTT GYRINSAKDD AAGLQISNRL SNQISGLNVA TRNANDGISL AQTAEGALQQ STNILQRIR DLALQSANGS NSDADRAALQ KEVAAQQAEL TRISDTTTFG GRKLLDGSFG TTSFQVGSNA YETIDISLQN A SASAIGSY ...String: VNTNIASLNT QRNLNASSND LNTSLQRLTT GYRINSAKDD AAGLQISNRL SNQISGLNVA TRNANDGISL AQTAEGALQQ STNILQRIR DLALQSANGS NSDADRAALQ KEVAAQQAEL TRISDTTTFG GRKLLDGSFG TTSFQVGSNA YETIDISLQN A SASAIGSY QVGSNGAGTV ASVAGTATAS GIASGTVNLV GGGQVKNIAI AAGDSAKAIA EKMDGAIPNL SARARTVFTA DV SGVTGGS LNFDVTVGSN TVSLAGVTST QDLADQLNSN SSKLGITASI NDKGVLTITS ATGENVKFGA QTGTATAGQV AVK VQGSDG KFEAAAKNVV AAGTAATTTI VTGYVQLNSP TAYSVSGTGT QASQVFGNAS AAQKSSVASV DISTADGAQN AIAV VDNAL AAIDAQRADL AAVQNRFKNT IDNLTNISEN ATNARSRIKD TDFAAETAAL SKNQVLQQAG TAILAQANQL PQAVL SLLR UniProtKB: B-type flagellin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)