+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the CBC-ALYREF complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | mRNA nuclear export / RNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsnRNA export from nucleus / nuclear cap binding complex / RNA cap binding complex / histone mRNA metabolic process / positive regulation of RNA export from nucleus / mRNA metabolic process / positive regulation of mRNA 3'-end processing / cap-dependent translational initiation / Processing of Intronless Pre-mRNAs / RNA cap binding ...snRNA export from nucleus / nuclear cap binding complex / RNA cap binding complex / histone mRNA metabolic process / positive regulation of RNA export from nucleus / mRNA metabolic process / positive regulation of mRNA 3'-end processing / cap-dependent translational initiation / Processing of Intronless Pre-mRNAs / RNA cap binding / snRNA binding / miRNA-mediated post-transcriptional gene silencing / Nuclear RNA decay / primary miRNA processing / regulation of mRNA processing / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / Transport of the SLBP independent Mature mRNA / Transport of the SLBP Dependant Mature mRNA / regulatory ncRNA-mediated post-transcriptional gene silencing / alternative mRNA splicing, via spliceosome / RNA 7-methylguanosine cap binding / positive regulation of mRNA splicing, via spliceosome / Transport of Mature mRNA Derived from an Intronless Transcript / mRNA 3'-end processing / mRNA cis splicing, via spliceosome / mRNA 3'-end processing / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA catabolic process / regulation of translational initiation / Abortive elongation of HIV-1 transcript in the absence of Tat / RNA Polymerase II Transcription Termination / FGFR2 alternative splicing / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / Signaling by FGFR2 IIIa TM / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / mRNA Splicing - Minor Pathway / spliceosomal complex assembly / Processing of Capped Intron-Containing Pre-mRNA / RNA polymerase II transcribes snRNA genes / 7-methylguanosine mRNA capping / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / mRNA transport / Formation of HIV-1 elongation complex containing HIV-1 Tat / mRNA export from nucleus / Formation of HIV elongation complex in the absence of HIV Tat / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Formation of RNA Pol II elongation complex / RNA Polymerase II Pre-transcription Events / mRNA Splicing - Major Pathway / RNA splicing / positive regulation of transcription elongation by RNA polymerase II / mRNA splicing, via spliceosome / mRNA transcription by RNA polymerase II / Regulation of expression of SLITs and ROBOs / single-stranded DNA binding / positive regulation of cell growth / snRNP Assembly / defense response to virus / molecular adaptor activity / ciliary basal body / ribonucleoprotein complex / mRNA binding / mitochondrion / DNA binding / RNA binding / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
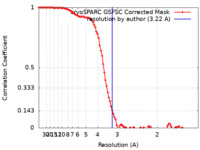
| Method | single particle reconstruction / cryo EM / Resolution: 3.22 Å | |||||||||
 Authors Authors | Xie Y / Clarke BP / Ren Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2024 Journal: Elife / Year: 2024Title: Cryo-EM structure of the CBC-ALYREF complex. Authors: Bradley P Clarke / Alexia E Angelos / Menghan Mei / Pate S Hill / Yihu Xie / Yi Ren /  Abstract: In eukaryotes, RNAs transcribed by RNA Pol II are modified at the 5' end with a 7-methylguanosine (mG) cap, which is recognized by the nuclear cap binding complex (CBC). The CBC plays multiple ...In eukaryotes, RNAs transcribed by RNA Pol II are modified at the 5' end with a 7-methylguanosine (mG) cap, which is recognized by the nuclear cap binding complex (CBC). The CBC plays multiple important roles in mRNA metabolism, including transcription, splicing, polyadenylation, and export. It promotes mRNA export through direct interaction with a key mRNA export factor, ALYREF, which in turn links the TRanscription and EXport (TREX) complex to the 5' end of mRNA. However, the molecular mechanism for CBC-mediated recruitment of the mRNA export machinery is not well understood. Here, we present the first structure of the CBC in complex with an mRNA export factor, ALYREF. The cryo-EM structure of CBC-ALYREF reveals that the RRM domain of ALYREF makes direct contact with both the NCBP1 and NCBP2 subunits of the CBC. Comparing CBC-ALYREF with other cellular complexes containing CBC and/or ALYREF components provides insights into the coordinated events during mRNA transcription, splicing, and export. #1:  Journal: Elife / Year: 2024 Journal: Elife / Year: 2024Title: Cryo-EM structure of the CBC-ALYREF complex Authors: Clarke BP / Angelos AE / Mei M / Hill PS / Xie Y / Ren Y | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40739.map.gz emd_40739.map.gz | 45.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40739-v30.xml emd-40739-v30.xml emd-40739.xml emd-40739.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40739_fsc.xml emd_40739_fsc.xml | 9.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_40739.png emd_40739.png | 81.7 KB | ||
| Filedesc metadata |  emd-40739.cif.gz emd-40739.cif.gz | 6 KB | ||
| Others |  emd_40739_half_map_1.map.gz emd_40739_half_map_1.map.gz emd_40739_half_map_2.map.gz emd_40739_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40739 http://ftp.pdbj.org/pub/emdb/structures/EMD-40739 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40739 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40739 | HTTPS FTP |
-Related structure data
| Related structure data |  8srrM M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40739.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40739.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.732 Å | ||||||||||||||||||||||||||||||||||||
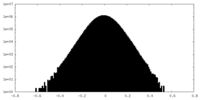
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40739_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
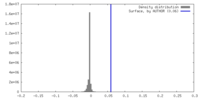
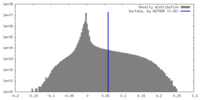
| Density Histograms |
-Half map: #1
| File | emd_40739_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of an mRNA export factor
| Entire | Name: Cryo-EM structure of an mRNA export factor |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of an mRNA export factor
| Supramolecule | Name: Cryo-EM structure of an mRNA export factor / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Nuclear cap-binding protein subunit 1
| Macromolecule | Name: Nuclear cap-binding protein subunit 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 91.960297 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSRRRHSDEN DGGQPHKRRK TSDANETEDH LESLICKVGE KSACSLESNL EGLAGVLEAD LPNYKSKILR LLCTVARLLP EKLTIYTTL VGLLNARNYN FGGEFVEAMI RQLKESLKAN NYNEAVYLVR FLSDLVNCHV IAAPSMVAMF ENFVSVTQEE D VPQVRRDW ...String: MSRRRHSDEN DGGQPHKRRK TSDANETEDH LESLICKVGE KSACSLESNL EGLAGVLEAD LPNYKSKILR LLCTVARLLP EKLTIYTTL VGLLNARNYN FGGEFVEAMI RQLKESLKAN NYNEAVYLVR FLSDLVNCHV IAAPSMVAMF ENFVSVTQEE D VPQVRRDW YVYAFLSSLP WVGKELYEKK DAEMDRIFAN TESYLKRRQK THVPMLQVWT ADKPHPQEEY LDCLWAQIQK LK KDRWQER HILRPYLAFD SILCEALQHN LPPFTPPPHT EDSVYPMPRV IFRMFDYTDD PEGPVMPGSH SVERFVIEEN LHC IIKSHW KERKTCAAQL VSYPGKNKIP LNYHIVEVIF AELFQLPAPP HIDVMYTTLL IELCKLQPGS LPQVLAQATE MLYM RLDTM NTTCVDRFIN WFSHHLSNFQ FRWSWEDWSD CLSQDPESPK PKFVREVLEK CMRLSYHQRI LDIVPPTFSA LCPAN PTCI YKYGDESSNS LPGHSVALCL AVAFKSKATN DEIFSILKDV PNPNQDDDDD EGFSFNPLKI EVFVQTLLHL AAKSFS HSF SALAKFHEVF KTLAESDEGK LHVLRVMFEV WRNHPQMIAV LVDKMIRTQI VDCAAVANWI FSSELSRDFT RLFVWEI LH STIRKMNKHV LKIQKELEEA KEKLARQHKR RSDDDDRSSD RKDGVLEEQI ERLQEKVESA QSEQKNLFLV IFQRFIMI L TEHLVRCETD GTSVLTPWYK NCIERLQQIF LQHHQIIQQY MVTLENLLFT AELDPHILAV FQQFCALQA UniProtKB: Nuclear cap-binding protein subunit 1 |
-Macromolecule #2: Nuclear cap-binding protein subunit 2
| Macromolecule | Name: Nuclear cap-binding protein subunit 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.028131 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSGGLLKALR SDSYVELSQY RDQHFRGDNE EQEKLLKKSC TLYVGNLSFY TTEEQIYELF SKSGDIKKII MGLDKMKKTA CGFCFVEYY SRADAENAMR YINGTRLDDR IIRTDWDAGF KEGRQYGRGR SGGQVRDEYR QDYDAGRGGY GKLAQNQ UniProtKB: Nuclear cap-binding protein subunit 2 |
-Macromolecule #3: RNA and export factor binding protein 2
| Macromolecule | Name: RNA and export factor binding protein 2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.542857 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GAMGSMADKM DMSLDDIIKL NRNQRRVNRG GGPRRNRPAI ARGGRNRPAP YSRPKPLPDK WQHDLFDSGC GGGEGVETGA KLLVSNLDF GVSDADIQEL FAEFGTLKKA AVDYDRSGRS LGTADVHFER RADALKAMKQ YKGVPLDGRP MDIQLVTSQI D UniProtKB: RNA and export factor binding protein 2 |
-Macromolecule #4: 7N-METHYL-8-HYDROGUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: 7N-METHYL-8-HYDROGUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 1 / Formula: M7G |
|---|---|
| Molecular weight | Theoretical: 458.235 Da |
| Chemical component information | 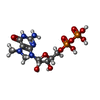 ChemComp-M7G: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)