+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Homo-octamer of PbuCsx28 protein | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR-associated protein / ANTIVIRAL PROTEIN | |||||||||
| Function / homology | Uncharacterized protein Function and homology information Function and homology information | |||||||||
| Biological species |  Prevotella buccae (bacteria) Prevotella buccae (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.65 Å | |||||||||
 Authors Authors | Park JU / Kellogg EH | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Csx28 is a membrane pore that enhances CRISPR-Cas13b-dependent antiphage defense. Authors: Arica R VanderWal / Jung-Un Park / Bogdan Polevoda / Julia K Nicosia / Adrian M Molina Vargas / Elizabeth H Kellogg / Mitchell R O'Connell /  Abstract: Type VI CRISPR-Cas systems use RNA-guided ribonuclease (RNase) Cas13 to defend bacteria against viruses, and some of these systems encode putative membrane proteins that have unclear roles in Cas13- ...Type VI CRISPR-Cas systems use RNA-guided ribonuclease (RNase) Cas13 to defend bacteria against viruses, and some of these systems encode putative membrane proteins that have unclear roles in Cas13-mediated defense. We show that Csx28, of type VI-B2 systems, is a transmembrane protein that assists to slow cellular metabolism upon viral infection, increasing antiviral defense. High-resolution cryo-electron microscopy reveals that Csx28 forms an octameric pore-like structure. These Csx28 pores localize to the inner membrane in vivo. Csx28's antiviral activity in vivo requires sequence-specific cleavage of viral messenger RNAs by Cas13b, which subsequently results in membrane depolarization, slowed metabolism, and inhibition of sustained viral infection. Our work suggests a mechanism by which Csx28 acts as a downstream, Cas13b-dependent effector protein that uses membrane perturbation as an antiviral defense strategy. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40059.map.gz emd_40059.map.gz | 3.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40059-v30.xml emd-40059-v30.xml emd-40059.xml emd-40059.xml | 17.6 KB 17.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40059_fsc.xml emd_40059_fsc.xml | 7.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_40059.png emd_40059.png | 176.5 KB | ||
| Filedesc metadata |  emd-40059.cif.gz emd-40059.cif.gz | 5.9 KB | ||
| Others |  emd_40059_half_map_1.map.gz emd_40059_half_map_1.map.gz emd_40059_half_map_2.map.gz emd_40059_half_map_2.map.gz | 27.9 MB 27.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40059 http://ftp.pdbj.org/pub/emdb/structures/EMD-40059 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40059 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40059 | HTTPS FTP |
-Related structure data
| Related structure data |  8gi1MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40059.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40059.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: halfmap 1
| File | emd_40059_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
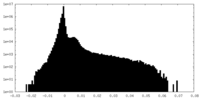
| Density Histograms |
-Half map: halfmap 2
| File | emd_40059_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
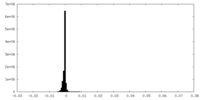
| Density Histograms |
- Sample components
Sample components
-Entire : octameric assembly of csx28
| Entire | Name: octameric assembly of csx28 |
|---|---|
| Components |
|
-Supramolecule #1: octameric assembly of csx28
| Supramolecule | Name: octameric assembly of csx28 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Prevotella buccae (bacteria) Prevotella buccae (bacteria) |
| Molecular weight | Theoretical: 300 KDa |
-Macromolecule #1: Accessory protein Csx28
| Macromolecule | Name: Accessory protein Csx28 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Prevotella buccae (bacteria) Prevotella buccae (bacteria) |
| Molecular weight | Theoretical: 21.910193 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDYMELAKEA FSIICTFIAA YVAYYYAIKQ LHQKSVENIE YAKYQAVLQA HKSLYKLLRF TTNTENEDSI LIWEKTKDGK QEATYYFRK ENIRKFIKEL SKEIYNEGCG IFMSKEALSL ISEYRNIVYG FMLSAQNNPQ ETIRITNRES VERMKKIHQN L SIEIRQAI NLKKRDLRFE NLYFQ UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.35 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: GRAPHENE OXIDE / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 47.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8gi1: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)