+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human apo GPR156 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / class C / GPR156 / cryo-EM / homodimer / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationstereocilium bundle organization / G protein-coupled receptor heterodimeric complex / G protein-coupled GABA receptor activity / gamma-aminobutyric acid signaling pathway / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.09 Å | |||||||||
 Authors Authors | Ma XY / Chen LN / Liao MH / Zhang LY / Xi K / Guo JM | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Molecular insights into the activation mechanism of GPR156 in maintaining auditory function. Authors: Xiangyu Ma / Li-Nan Chen / Menghui Liao / Liyan Zhang / Kun Xi / Jiamin Guo / Cangsong Shen / Dan-Dan Shen / Pengjun Cai / Qingya Shen / Jieyu Qi / Huibing Zhang / Shao-Kun Zang / Ying-Jun ...Authors: Xiangyu Ma / Li-Nan Chen / Menghui Liao / Liyan Zhang / Kun Xi / Jiamin Guo / Cangsong Shen / Dan-Dan Shen / Pengjun Cai / Qingya Shen / Jieyu Qi / Huibing Zhang / Shao-Kun Zang / Ying-Jun Dong / Luwei Miao / Jiao Qin / Su-Yu Ji / Yue Li / Jianfeng Liu / Chunyou Mao / Yan Zhang / Renjie Chai /  Abstract: The class C orphan G-protein-coupled receptor (GPCR) GPR156, which lacks the large extracellular region, plays a pivotal role in auditory function through G. Here, we firstly demonstrate that GPR156 ...The class C orphan G-protein-coupled receptor (GPCR) GPR156, which lacks the large extracellular region, plays a pivotal role in auditory function through G. Here, we firstly demonstrate that GPR156 with high constitutive activity is essential for maintaining auditory function, and further reveal the structural basis of the sustained role of GPR156. We present the cryo-EM structures of human apo GPR156 and the GPR156-G complex, unveiling a small extracellular region formed by extracellular loop 2 (ECL2) and the N-terminus. The GPR156 dimer in both apo state and G protein-coupled state adopt a transmembrane (TM)5/6-TM5/6 interface, indicating the high constitutive activity of GPR156 in the apo state. Furthermore, C-terminus in G-bound subunit of GPR156 plays a dual role in promoting G protein binding within G-bound subunit while preventing the G-free subunit from binding to additional G protein. Together, these results explain how GPR156 constitutive activity is maintained through dimerization and provide a mechanistic insight into the sustained role of GPR156 in maintaining auditory function. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39345.map.gz emd_39345.map.gz | 56.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39345-v30.xml emd-39345-v30.xml emd-39345.xml emd-39345.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_39345.png emd_39345.png | 103.8 KB | ||
| Filedesc metadata |  emd-39345.cif.gz emd-39345.cif.gz | 6 KB | ||
| Others |  emd_39345_half_map_1.map.gz emd_39345_half_map_1.map.gz emd_39345_half_map_2.map.gz emd_39345_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39345 http://ftp.pdbj.org/pub/emdb/structures/EMD-39345 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39345 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39345 | HTTPS FTP |
-Related structure data
| Related structure data |  8yjpMC  8yk0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_39345.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39345.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.93 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_39345_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_39345_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GPR156
| Entire | Name: GPR156 |
|---|---|
| Components |
|
-Supramolecule #1: GPR156
| Supramolecule | Name: GPR156 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Probable G-protein coupled receptor 156
| Macromolecule | Name: Probable G-protein coupled receptor 156 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.47223 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEPEINCSEL CDSFPGQELD RRPLHDLCKT TITSSHHSSK TISSLSPVLL GIVWTFLSCG LLLILFFLAF TIHCRKNRIV KMSSPNLNI VTLLGSCLTY SSAYLFGIQD VLVGSSMETL IQTRLSMLCI GTSLVFGPIL GKSWRLYKVF TQRVPDKRVI I KDLQLLGL ...String: MEPEINCSEL CDSFPGQELD RRPLHDLCKT TITSSHHSSK TISSLSPVLL GIVWTFLSCG LLLILFFLAF TIHCRKNRIV KMSSPNLNI VTLLGSCLTY SSAYLFGIQD VLVGSSMETL IQTRLSMLCI GTSLVFGPIL GKSWRLYKVF TQRVPDKRVI I KDLQLLGL VAALLMADVI LLMTWVLTDP IQCLQILSVS MTVTGKDVSC TSTSTHFCAS RYSDVWIALI WGCKGLLLLY GA YLAGLTG HVSSPPVNQS LTIMVGVNLL VLAAGLLFVV TRYLHSWPNL VFGLTSGGIF VCTTTINCFI FIPQLKQWKA FEE ENQTIR RMAKYFSTPN KSFHTQYGEE UniProtKB: Probable G-protein coupled receptor 156 |
-Macromolecule #2: (21R,24R,27S)-24,27,28-trihydroxy-18,24-dioxo-19,23,25-trioxa-24l...
| Macromolecule | Name: (21R,24R,27S)-24,27,28-trihydroxy-18,24-dioxo-19,23,25-trioxa-24lambda~5~-phosphaoctacosan-21-yl (9Z)-octadec-9-enoate type: ligand / ID: 2 / Number of copies: 4 / Formula: MW9 |
|---|---|
| Molecular weight | Theoretical: 777.06 Da |
| Chemical component information | 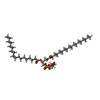 ChemComp-MW9: |
-Macromolecule #3: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 3 / Number of copies: 9 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































