[English] 日本語
 Yorodumi
Yorodumi- EMDB-39180: Non-catalytic site depleted and epsilon C-terminal domain deleted... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Non-catalytic site depleted and epsilon C-terminal domain deleted FoF1-ATPase from Bacillus PS3,state2,nucleotide depleted condition | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / MEMBRANE PROTEIN | |||||||||
| Biological species |  | |||||||||
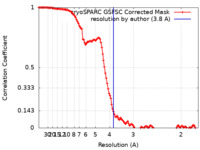
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Kobayashi R / Nakano A / Mitsuoka K / Yokoyoma K | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Biochim Biophys Acta Bioenerg / Year: 2025 Journal: Biochim Biophys Acta Bioenerg / Year: 2025Title: ADP-inhibited structure of non-catalytic site-depleted FF-ATPase from thermophilic Bacillus sp. PS-3. Authors: Ren Kobayashi / Astuki Nakano / Kaoru Mitsuoka / Ken Yokoyama /  Abstract: The F domain of FF-ATP synthases/ATPases (FF) possesses three catalytic sites on the three αβ interfaces, termed αβ, αβ, and αβ, located mainly on the β subunits. The enzyme also has three ...The F domain of FF-ATP synthases/ATPases (FF) possesses three catalytic sites on the three αβ interfaces, termed αβ, αβ, and αβ, located mainly on the β subunits. The enzyme also has three non-catalytic ATP-binding sites on the three αβ interfaces, located mainly on the α subunits. When ATP does not bind to the non-catalytic site, FF becomes significantly prone to ADP inhibition, ultimately resulting in the loss of ATPase activity. However, the underlying mechanism of ADP inhibition remains unclear. Here, we report the cryo-EM structure of the non-catalytic site-depleted (ΔNC) FF from thermophilic Bacillus sp. PS-3, which completely lacks the ability to bind ATP (and ADP) upon transitioning to the ADP-inhibited form. The structure closely resembled the 81° rotated structure of the wild-type FF, except for minor movements in the C-terminal region of the α subunit. In this structure, unlike the wild-type enzyme, the catalytic site at αβ, responsible for ATP hydrolysis, was occupied by ADP-Mg, with the absence of Pi. Furthermore, the catalytic site at αβ, where ATP enters the F domain during steady-state catalysis, is occupied by ADP, seemingly impeding further ATP binding to the enzyme. The structure suggests that the ADP-inhibited form of the F domain is more likely due to differences in the nucleotide-binding states at the catalytic sites rather than structural differences. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39180.map.gz emd_39180.map.gz | 88.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39180-v30.xml emd-39180-v30.xml emd-39180.xml emd-39180.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39180_fsc.xml emd_39180_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_39180.png emd_39180.png | 106.5 KB | ||
| Masks |  emd_39180_msk_1.map emd_39180_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-39180.cif.gz emd-39180.cif.gz | 3.9 KB | ||
| Others |  emd_39180_additional_1.map.gz emd_39180_additional_1.map.gz emd_39180_additional_2.map.gz emd_39180_additional_2.map.gz emd_39180_half_map_1.map.gz emd_39180_half_map_1.map.gz emd_39180_half_map_2.map.gz emd_39180_half_map_2.map.gz | 168.1 MB 12.1 MB 165 MB 165 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39180 http://ftp.pdbj.org/pub/emdb/structures/EMD-39180 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39180 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39180 | HTTPS FTP |
-Validation report
| Summary document |  emd_39180_validation.pdf.gz emd_39180_validation.pdf.gz | 1.3 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_39180_full_validation.pdf.gz emd_39180_full_validation.pdf.gz | 1.3 MB | Display | |
| Data in XML |  emd_39180_validation.xml.gz emd_39180_validation.xml.gz | 20.6 KB | Display | |
| Data in CIF |  emd_39180_validation.cif.gz emd_39180_validation.cif.gz | 26.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39180 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39180 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39180 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39180 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_39180.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39180.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_39180_msk_1.map emd_39180_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_39180_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_39180_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_39180_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_39180_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FoF1 from Bacillus sp. PS3
| Entire | Name: FoF1 from Bacillus sp. PS3 |
|---|---|
| Components |
|
-Supramolecule #1: FoF1 from Bacillus sp. PS3
| Supramolecule | Name: FoF1 from Bacillus sp. PS3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)