[English] 日本語
 Yorodumi
Yorodumi- EMDB-3860: Cryo-EM map of calcium-bound mTMEM16A chloride channel at 3.75 A ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3860 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of calcium-bound mTMEM16A chloride channel at 3.75 A resolution | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TMEM16 family / ion channel / membrane protein / cryo-EM | |||||||||
| Function / homology |  Function and homology information Function and homology informationglial cell projection elongation / trachea development / iodide transmembrane transporter activity / mucus secretion / iodide transport / intracellularly calcium-gated chloride channel activity / cellular response to peptide / Stimuli-sensing channels / voltage-gated chloride channel activity / calcium-activated cation channel activity ...glial cell projection elongation / trachea development / iodide transmembrane transporter activity / mucus secretion / iodide transport / intracellularly calcium-gated chloride channel activity / cellular response to peptide / Stimuli-sensing channels / voltage-gated chloride channel activity / calcium-activated cation channel activity / protein localization to membrane / chloride transport / detection of temperature stimulus involved in sensory perception of pain / chloride channel activity / chloride channel complex / chloride transmembrane transport / regulation of membrane potential / cellular response to heat / presynaptic membrane / phospholipase C-activating G protein-coupled receptor signaling pathway / apical plasma membrane / signaling receptor binding / external side of plasma membrane / glutamatergic synapse / protein homodimerization activity / nucleoplasm / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | |||||||||
 Authors Authors | Paulino C / Kalienkova V | |||||||||
| Funding support |  Switzerland, 2 items Switzerland, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Activation mechanism of the calcium-activated chloride channel TMEM16A revealed by cryo-EM. Authors: Cristina Paulino / Valeria Kalienkova / Andy K M Lam / Yvonne Neldner / Raimund Dutzler /  Abstract: The calcium-activated chloride channel TMEM16A is a ligand-gated anion channel that opens in response to an increase in intracellular Ca concentration. The protein is broadly expressed and ...The calcium-activated chloride channel TMEM16A is a ligand-gated anion channel that opens in response to an increase in intracellular Ca concentration. The protein is broadly expressed and contributes to diverse physiological processes, including transepithelial chloride transport and the control of electrical signalling in smooth muscles and certain neurons. As a member of the TMEM16 (or anoctamin) family of membrane proteins, TMEM16A is closely related to paralogues that function as scramblases, which facilitate the bidirectional movement of lipids across membranes. The unusual functional diversity of the TMEM16 family and the relationship between two seemingly incompatible transport mechanisms has been the focus of recent investigations. Previous breakthroughs were obtained from the X-ray structure of the lipid scramblase of the fungus Nectria haematococca (nhTMEM16), and from the cryo-electron microscopy structure of mouse TMEM16A at 6.6 Å (ref. 14). Although the latter structure disclosed the architectural differences that distinguish ion channels from lipid scramblases, its low resolution did not permit a detailed molecular description of the protein or provide any insight into its activation by Ca. Here we describe the structures of mouse TMEM16A at high resolution in the presence and absence of Ca. These structures reveal the differences between ligand-bound and ligand-free states of a calcium-activated chloride channel, and when combined with functional experiments suggest a mechanism for gating. During activation, the binding of Ca to a site located within the transmembrane domain, in the vicinity of the pore, alters the electrostatic properties of the ion conduction path and triggers a conformational rearrangement of an α-helix that comes into physical contact with the bound ligand, and thereby directly couples ligand binding and pore opening. Our study describes a process that is unique among channel proteins, but one that is presumably general for both functional branches of the TMEM16 family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3860.map.gz emd_3860.map.gz | 96.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3860-v30.xml emd-3860-v30.xml emd-3860.xml emd-3860.xml | 24.9 KB 24.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3860_fsc.xml emd_3860_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_3860.png emd_3860.png | 235 KB | ||
| Masks |  emd_3860_msk_1.map emd_3860_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-3860.cif.gz emd-3860.cif.gz | 7.9 KB | ||
| Others |  emd_3860_additional.map.gz emd_3860_additional.map.gz emd_3860_half_map_1.map.gz emd_3860_half_map_1.map.gz emd_3860_half_map_2.map.gz emd_3860_half_map_2.map.gz | 96.4 MB 96.2 MB 96.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3860 http://ftp.pdbj.org/pub/emdb/structures/EMD-3860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3860 | HTTPS FTP |
-Related structure data
| Related structure data |  5oybMC  3861C  5oygC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3860.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3860.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.075 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_3860_msk_1.map emd_3860_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
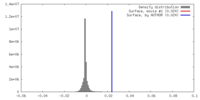
| Density Histograms |
-Additional map: None
| File | emd_3860_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
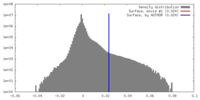
| Density Histograms |
-Half map: cryo-EM half map 2 of the ion channel...
| File | emd_3860_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map 2 of the ion channel TMEM16A from mouse in presence of calcium ions | ||||||||||||
| Projections & Slices |
| ||||||||||||
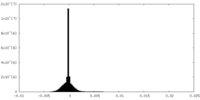
| Density Histograms |
-Half map: cryo-EM half map 1 of the ion channel...
| File | emd_3860_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map 1 of the ion channel TMEM16A from mouse in presence of calcium ions | ||||||||||||
| Projections & Slices |
| ||||||||||||
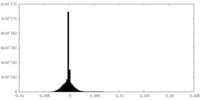
| Density Histograms |
- Sample components
Sample components
-Entire : mTMEM16A with calcium ions bound
| Entire | Name: mTMEM16A with calcium ions bound |
|---|---|
| Components |
|
-Supramolecule #1: mTMEM16A with calcium ions bound
| Supramolecule | Name: mTMEM16A with calcium ions bound / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: calcium-activated chloride channel |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 110.916 KDa |
-Macromolecule #1: Anoctamin-1
| Macromolecule | Name: Anoctamin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 111.058992 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRVPEKYSTL PAEDRSVHIV NICAIEDLGY LPSEGTLLNS LSVDPDAECK YGLYFRDGKR KVDYILVYHH KRASGSRTLA RRGLQNDMV LGTRSVRQDQ PLPGKGSPVD AGSPEVPMDY HEDDKRFRRE EYEGNLLEAG LELENDEDTK IHGVGFVKIH A PWHVLCRE ...String: MRVPEKYSTL PAEDRSVHIV NICAIEDLGY LPSEGTLLNS LSVDPDAECK YGLYFRDGKR KVDYILVYHH KRASGSRTLA RRGLQNDMV LGTRSVRQDQ PLPGKGSPVD AGSPEVPMDY HEDDKRFRRE EYEGNLLEAG LELENDEDTK IHGVGFVKIH A PWHVLCRE AEFLKLKMPT KKVYHISETR GLLKTINSVL QKITDPIQPK VAEHRPQTTK RLSYPFSREK QHLFDLTDRD SF FDSKTRS TIVYEILKRT TCTKAKYSMG ITSLLANGVY SAAYPLHDGD YEGDNVEFND RKLLYEEWAS YGVFYKYQPI DLV RKYFGE KVGLYFAWLG AYTQMLIPAS IVGVIVFLYG CATVDENIPS MEMCDQRYNI TMCPLCDKTC SYWKMSSACA TARA SHLFD NPATVFFSVF MALWAATFME HWKRKQMRLN YRWDLTGFEE EEEAVKDHPR AEYEARVLEK SLRKESRNKE TDKVK LTWR DRFPAYFTNL VSIIFMIAVT FAIVLGVIIY RISTAAALAM NSSPSVRSNI RVTVTATAVI INLVVIILLD EVYGCI ARW LTKIEVPKTE KSFEERLTFK AFLLKFVNSY TPIFYVAFFK GRFVGRPGDY VYIFRSFRME ECAPGGCLME LCIQLSI IM LGKQLIQNNL FEIGIPKMKK FIRYLKLRRQ SPSDREEYVK RKQRYEVDFN LEPFAGLTPE YMEMIIQFGF VTLFVASF P LAPLFALLNN IIEIRLDAKK FVTELRRPVA IRAKDIGIWY NILRGVGKLA VIINAFVISF TSDFIPRLVY LYMYSQNGT MHGFVNHTLS SFNVSDFQNG TAPNDPLDLG YEVQICRYKD YREPPWSEHK YDISKDFWAV LAARLAFVIV FQNLVMFMSD FVDWVIPDI PKDISQQIHK EKVLMVELFM REEQGKQQLL DTWMEKEKPR DVPCNNHSPT THPEAGDGSP VPSYEYHGDA L UniProtKB: Anoctamin-1 |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Component - Concentration: 20.0 mM / Component - Name: Hepes / Details: 20 mM Hepes 150 mM NaCl 0.5 mM CaCl2 <0.12% digitonin |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 288 K / Instrument: FEI VITROBOT MARK IV / Details: 2 ul sample volume 2-4 sec blotting time. |
| Details | full-length (wild-type isoform ac) deglycosylated mTMEM16A in presence of 0.5mM CaCl2 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80.0 K / Max: 100.0 K |
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Lower energy threshold: -10 eV / Energy filter - Upper energy threshold: +10 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 7420 pixel / Digitization - Dimensions - Height: 7676 pixel / Digitization - Frames/image: 1-80 / Number grids imaged: 3 / Number real images: 4343 / Average exposure time: 10.0 sec. / Average electron dose: 80.0 e/Å2 Details: Data were collected in an automated fashion using SerialEM47 on a K2 Summit detector (Gatan). For the dataset in presence of calcium ions, cryo-EM images were collected at a pixel size of 0. ...Details: Data were collected in an automated fashion using SerialEM47 on a K2 Summit detector (Gatan). For the dataset in presence of calcium ions, cryo-EM images were collected at a pixel size of 0.5375A in super-resolution mode, a defocus range of -0.5 to -3.0 um, an exposure time of 10 sec and a sub-frame exposure time of 125 ms (80 frames) with an approximate electron dose of 1 e-/A2/frame. An extensive effort was made for this dataset to screen different areas within the grid and only regions that provided an estimated resolution of the CTF fit of better than 4A were selected for data collection. The total accumulated dose on the specimen level was approximately 80 e-/A2. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.5 µm / Calibrated magnification: 46511 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 46511 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-5oyb: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)