+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
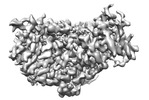
| Title | local refinement of MCM_ATP_dsDNA | |||||||||
 Map data Map data | local refinement of MCM_ATP_dsDNA | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Helicase / Replication / HYDROLASE | |||||||||
| Biological species |   Thermococcus kodakarensis (archaea) Thermococcus kodakarensis (archaea) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.57 Å | |||||||||
 Authors Authors | Ma J / Yi G / Ye M / MacGregor-Chatwin C / Sheng Y / Lu Y / Li M / Gilbert RJC / Zhang P | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Open architecture of archaea MCM and dsDNA complexes resolved using monodispersed streptavidin affinity CryoEM. Authors: Jianbing Ma / Gangshun Yi / Mingda Ye / Craig MacGregor-Chatwin / Yuewen Sheng / Ying Lu / Ming Li / Qingrong Li / Dong Wang / Robert J C Gilbert / Peijun Zhang /    Abstract: The cryo-electron microscopy (cryoEM) method has enabled high-resolution structure determination of numerous biomolecules and complexes. Nevertheless, cryoEM sample preparation of challenging ...The cryo-electron microscopy (cryoEM) method has enabled high-resolution structure determination of numerous biomolecules and complexes. Nevertheless, cryoEM sample preparation of challenging proteins and complexes, especially those with low abundance or with preferential orientation, remains a major hurdle. We developed an affinity-grid method employing monodispersed single particle streptavidin on a lipid monolayer to enhance particle absorption on the grid surface and alleviate sample exposure to the air-water interface. Using this approach, we successfully enriched the Thermococcus kodakarensis mini-chromosome maintenance complex 3 (MCM3) on cryoEM grids through biotinylation and resolved its structure. We further utilized this affinity method to tether the biotin-tagged dsDNA to selectively enrich a stable MCM3-ATP-dsDNA complex for cryoEM structure determination. Intriguingly, both MCM3 apo and dsDNA bound structures exhibit left-handed open spiral conformations, distinct from other reported MCM structures. The large open gate is sufficient to accommodate a dsDNA which could potentially be melted. The value of mspSA affinity method was further demonstrated by mitigating the issue of preferential angular distribution of HIV-1 capsid protein hexamer and RNA polymerase II elongation complex from Saccharomyces cerevisiae. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38123.map.gz emd_38123.map.gz | 11.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38123-v30.xml emd-38123-v30.xml emd-38123.xml emd-38123.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38123.png emd_38123.png | 79.4 KB | ||
| Filedesc metadata |  emd-38123.cif.gz emd-38123.cif.gz | 4 KB | ||
| Others |  emd_38123_half_map_1.map.gz emd_38123_half_map_1.map.gz emd_38123_half_map_2.map.gz emd_38123_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38123 http://ftp.pdbj.org/pub/emdb/structures/EMD-38123 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38123 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38123 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38123.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38123.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | local refinement of MCM_ATP_dsDNA | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.932 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map A of local refinement of MCM ATP dsDNA
| File | emd_38123_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A of local refinement of MCM_ATP_dsDNA | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B of local refinement of MCM ATP dsDNA
| File | emd_38123_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B of local refinement of MCM_ATP_dsDNA | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : MCM homohexamer in complex with dsDNA in presence of ATP
| Entire | Name: MCM homohexamer in complex with dsDNA in presence of ATP |
|---|---|
| Components |
|
-Supramolecule #1: MCM homohexamer in complex with dsDNA in presence of ATP
| Supramolecule | Name: MCM homohexamer in complex with dsDNA in presence of ATP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Thermococcus kodakarensis (archaea) Thermococcus kodakarensis (archaea) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER Details: Generated using ab-initio reconstruction routine in cryoSPARC |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.57 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 126808 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)