+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | One RBD up state of Spike glycoprotein, SARS-CoV-2 | |||||||||
 Map data Map data | Sharpened full map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Glycoprotein / Viral protein / SARS-CoV-2 | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
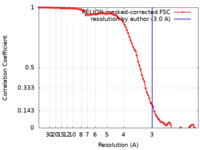
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Yadav S / Vinothkumar KR | |||||||||
| Funding support |  India, 2 items India, 2 items
| |||||||||
 Citation Citation |  Journal: Acta Crystallogr D Struct Biol / Year: 2024 Journal: Acta Crystallogr D Struct Biol / Year: 2024Title: Factors affecting macromolecule orientations in thin films formed in cryo-EM. Authors: Swati Yadav / Kutti R Vinothkumar /  Abstract: The formation of a vitrified thin film embedded with randomly oriented macromolecules is an essential prerequisite for cryogenic sample electron microscopy. Most commonly, this is achieved using the ...The formation of a vitrified thin film embedded with randomly oriented macromolecules is an essential prerequisite for cryogenic sample electron microscopy. Most commonly, this is achieved using the plunge-freeze method first described nearly 40 years ago. Although this is a robust method, the behaviour of different macromolecules shows great variation upon freezing and often needs to be optimized to obtain an isotropic, high-resolution reconstruction. For a macromolecule in such a film, the probability of encountering the air-water interface in the time between blotting and freezing and adopting preferred orientations is very high. 3D reconstruction using preferentially oriented particles often leads to anisotropic and uninterpretable maps. Currently, there are no general solutions to this prevalent issue, but several approaches largely focusing on sample preparation with the use of additives and novel grid modifications have been attempted. In this study, the effect of physical and chemical factors on the orientations of macromolecules was investigated through an analysis of selected well studied macromolecules, and important parameters that determine the behaviour of proteins on cryo-EM grids were revealed. These insights highlight the nature of the interactions that cause preferred orientations and can be utilized to systematically address orientation bias for any given macromolecule and to provide a framework to design small-molecule additives to enhance sample stability and behaviour. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37953.map.gz emd_37953.map.gz | 58.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37953-v30.xml emd-37953-v30.xml emd-37953.xml emd-37953.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37953_fsc.xml emd_37953_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_37953.png emd_37953.png | 20.7 KB | ||
| Filedesc metadata |  emd-37953.cif.gz emd-37953.cif.gz | 7.3 KB | ||
| Others |  emd_37953_half_map_1.map.gz emd_37953_half_map_1.map.gz emd_37953_half_map_2.map.gz emd_37953_half_map_2.map.gz | 49.5 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37953 http://ftp.pdbj.org/pub/emdb/structures/EMD-37953 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37953 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37953 | HTTPS FTP |
-Related structure data
| Related structure data |  8wziMC  8wv4C  8wv5C  8wv6C  8wzhC  8wzjC  8wzkC  8wzmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37953.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37953.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened full map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Unsharpened half map
| File | emd_37953_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half map | ||||||||||||
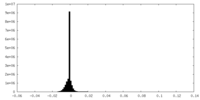
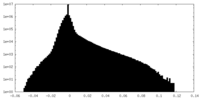
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unsharpened half map
| File | emd_37953_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half map | ||||||||||||
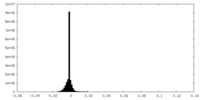
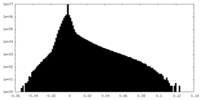
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Spike glycoprotein, SARS-CoV-2
| Entire | Name: Spike glycoprotein, SARS-CoV-2 |
|---|---|
| Components |
|
-Supramolecule #1: Spike glycoprotein, SARS-CoV-2
| Supramolecule | Name: Spike glycoprotein, SARS-CoV-2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 432 KDa |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 Details: linker (1216-SGRLVPRGSP-1225) and a fibritin trimerization motif (1226-GSGYIPEAPRDGQAYVRKDGEWVLLSTFLG-1254) followed by another linker (1256-GTK-1258) followed by the HRV3c post cleavage ...Details: linker (1216-SGRLVPRGSP-1225) and a fibritin trimerization motif (1226-GSGYIPEAPRDGQAYVRKDGEWVLLSTFLG-1254) followed by another linker (1256-GTK-1258) followed by the HRV3c post cleavage fragment (1259-LEVLFQ-1264) Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 139.706984 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAMFVFLVLL PLVSSQCVNL TTRTQLPPAY TNSFTRGVYY PDKVFRSSVL HSTQDLFLPF FSNVTWFHAI HVSGTNGTKR FDNPVLPFN DGVYFASTEK SNIIRGWIFG TTLDSKTQSL LIVNNATNVV IKVCEFQFCN DPFLGVYYHK NNKSWMESEF R VYSSANNC ...String: MAMFVFLVLL PLVSSQCVNL TTRTQLPPAY TNSFTRGVYY PDKVFRSSVL HSTQDLFLPF FSNVTWFHAI HVSGTNGTKR FDNPVLPFN DGVYFASTEK SNIIRGWIFG TTLDSKTQSL LIVNNATNVV IKVCEFQFCN DPFLGVYYHK NNKSWMESEF R VYSSANNC TFEYVSQPFL MDLEGKQGNF KNLREFVFKN IDGYFKIYSK HTPINLVRDL PQGFSALEPL VDLPIGINIT RF QTLLALH RSYLTPGDSS SGWTAGAAAY YVGYLQPRTF LLKYNENGTI TDAVDCALDP LSETKCTLKS FTVEKGIYQT SNF RVQPTE SIVRFPNITN LCPFGEVFNA TRFASVYAWN RKRISNCVAD YSVLYNSASF STFKCYGVSP TKLNDLCFTN VYAD SFVIR GDEVRQIAPG QTGKIADYNY KLPDDFTGCV IAWNSNNLDS KVGGNYNYLY RLFRKSNLKP FERDISTEIY QAGST PCNG VEGFNCYFPL QSYGFQPTNG VGYQPYRVVV LSFELLHAPA TVCGPKKSTN LVKNKCVNFN FNGLTGTGVL TESNKK FLP FQQFGRDIAD TTDAVRDPQT LEILDITPCS FGGVSVITPG TNTSNQVAVL YQDVNCTEVP VAIHADQLTP TWRVYST GS NVFQTRAGCL IGAEHVNNSY ECDIPIGAGI CASYQTQTNS PASVASQSII AYTMSLGAEN SVAYSNNSIA IPTNFTIS V TTEILPVSMT KTSVDCTMYI CGDSTECSNL LLQYGSFCTQ LNRALTGIAV EQDKNTQEVF AQVKQIYKTP PIKDFGGFN FSQILPDPSK PSKRSFIEDL LFNKVTLADA GFIKQYGDCL GDIAARDLIC AQKFNGLTVL PPLLTDEMIA QYTSALLAGT ITSGWTFGA GAALQIPFAM QMAYRFNGIG VTQNVLYENQ KLIANQFNSA IGKIQDSLSS TASALGKLQD VVNQNAQALN T LVKQLSSN FGAISSVLND ILSRLDPPEA EVQIDRLITG RLQSLQTYVT QQLIRAAEIR ASANLAATKM SECVLGQSKR VD FCGKGYH LMSFPQSAPH GVVFLHVTYV PAQEKNFTTA PAICHDGKAH FPREGVFVSN GTHWFVTQRN FYEPQIITTD NTF VSGNCD VVIGIVNNTV YDPLQPELDS FKEELDKYFK NHTSPDVDLG DISGINASVV NIQKEIDRLN EVAKNLNESL IDLQ ELGKY EQYIKWPSGR LVPRGSPGSG YIPEAPRDGQ AYVRKDGEWV LLSTFLGGTK LEVLFQ UniProtKB: Spike glycoprotein |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force, 0. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Average exposure time: 60.0 sec. / Average electron dose: 25.0 e/Å2 / Details: Movies were collected at 40 frames per second |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 130841 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)