[English] 日本語
 Yorodumi
Yorodumi- EMDB-37562: Cryo- EM structure of Mycobacterium smegmatis 70S ribosome, E- tR... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo- EM structure of Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH. | |||||||||
 Map data Map data | Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome / protein synthesis / Mycobacterium smegmatis / hibernation promotion factor / RafH / hypoxia stress / Cryo- EM / Single particle reconstruction. | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of translational elongation / ribosomal small subunit binding / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly ...negative regulation of translational elongation / ribosomal small subunit binding / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / RNA binding / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Kumar N / Sharma S / Kaushal PS | |||||||||
| Funding support |  India, 1 items India, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH. Authors: Niraj Kumar / Shivani Sharma / Prem S Kaushal /  Abstract: Ribosome hibernation is a key survival strategy bacteria adopt under environmental stress, where a protein, hibernation promotion factor (HPF), transitorily inactivates the ribosome. Mycobacterium ...Ribosome hibernation is a key survival strategy bacteria adopt under environmental stress, where a protein, hibernation promotion factor (HPF), transitorily inactivates the ribosome. Mycobacterium tuberculosis encounters hypoxia (low oxygen) as a major stress in the host macrophages, and upregulates the expression of RafH protein, which is crucial for its survival. The RafH, a dual domain HPF, an orthologue of bacterial long HPF (HPF), hibernates ribosome in 70S monosome form, whereas in other bacteria, the HPF induces 70S ribosome dimerization and hibernates its ribosome in 100S disome form. Here, we report the cryo- EM structure of M. smegmatis, a close homolog of M. tuberculosis, 70S ribosome in complex with the RafH factor at an overall 2.8 Å resolution. The N- terminus domain (NTD) of RafH binds to the decoding center, similarly to HPF NTD. In contrast, the C- terminus domain (CTD) of RafH, which is larger than the HPF CTD, binds to a distinct site at the platform binding center of the ribosomal small subunit. The two domain-connecting linker regions, which remain mostly disordered in earlier reported HPF structures, interact mainly with the anti-Shine Dalgarno sequence of the 16S rRNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37562.map.gz emd_37562.map.gz | 21.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37562-v30.xml emd-37562-v30.xml emd-37562.xml emd-37562.xml | 75.4 KB 75.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37562_fsc.xml emd_37562_fsc.xml | 13.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_37562.png emd_37562.png | 109 KB | ||
| Filedesc metadata |  emd-37562.cif.gz emd-37562.cif.gz | 13.8 KB | ||
| Others |  emd_37562_half_map_1.map.gz emd_37562_half_map_1.map.gz emd_37562_half_map_2.map.gz emd_37562_half_map_2.map.gz | 165.3 MB 165.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37562 http://ftp.pdbj.org/pub/emdb/structures/EMD-37562 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37562 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37562 | HTTPS FTP |
-Related structure data
| Related structure data |  8wibMC  8whxC  8whyC  8wi7C  8wi8C  8wi9C  8wicC  8widC  8wifC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37562.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37562.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
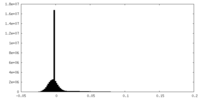
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH, half 2.
| File | emd_37562_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH, half 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH, half 1.
| File | emd_37562_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis 70S ribosome, E- tRNA and RafH, half 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : 70S ribosome + RafH protein
+Supramolecule #1: 70S ribosome + RafH protein
+Supramolecule #2: 70S ribosome
+Supramolecule #3: RafH
+Macromolecule #1: 50S ribosomal protein L2
+Macromolecule #2: 50S ribosomal protein L3
+Macromolecule #3: 50S ribosomal protein L4
+Macromolecule #4: 50S ribosomal protein L5
+Macromolecule #5: 50S ribosomal protein L6
+Macromolecule #6: 50S ribosomal protein L9
+Macromolecule #7: 50S ribosomal protein L13
+Macromolecule #8: 50S ribosomal protein L14
+Macromolecule #9: 50S ribosomal protein L15
+Macromolecule #10: 50S ribosomal protein L17
+Macromolecule #11: 50S ribosomal protein L18
+Macromolecule #12: 50S ribosomal protein L19
+Macromolecule #13: 50S ribosomal protein L20
+Macromolecule #14: 50S ribosomal protein L21
+Macromolecule #15: 50S ribosomal protein L22
+Macromolecule #16: 50S ribosomal protein L23
+Macromolecule #17: 50S ribosomal protein L24
+Macromolecule #18: 50S ribosomal protein L27
+Macromolecule #19: 50S ribosomal protein L28
+Macromolecule #20: 50S ribosomal protein L29
+Macromolecule #21: 50S ribosomal protein L30
+Macromolecule #22: 50S ribosomal protein L32
+Macromolecule #23: 50S ribosomal protein L33A
+Macromolecule #24: 50S ribosomal protein L34
+Macromolecule #25: 50S ribosomal protein L35
+Macromolecule #26: 50S ribosomal protein L31
+Macromolecule #31: 30S ribosomal protein S22
+Macromolecule #32: 30S ribosomal protein S3
+Macromolecule #33: 30S ribosomal protein S4
+Macromolecule #34: 30S ribosomal protein S5
+Macromolecule #35: 30S ribosomal protein S6
+Macromolecule #36: 30S ribosomal protein S7
+Macromolecule #37: 30S ribosomal protein S8
+Macromolecule #38: 30S ribosomal protein S9
+Macromolecule #39: 30S ribosomal protein S10
+Macromolecule #40: 30S ribosomal protein S11
+Macromolecule #41: 30S ribosomal protein S12
+Macromolecule #42: 30S ribosomal protein S13
+Macromolecule #43: 30S ribosomal protein S14A
+Macromolecule #44: 30S ribosomal protein S15
+Macromolecule #45: 30S ribosomal protein S16
+Macromolecule #46: 30S ribosomal protein S17
+Macromolecule #47: 30S ribosomal protein S18B
+Macromolecule #48: 30S ribosomal protein S19
+Macromolecule #49: 30S ribosomal protein S20
+Macromolecule #50: Ribosome hibernation promotion factor RafH
+Macromolecule #27: 23S rRNA
+Macromolecule #28: 5S rRNA
+Macromolecule #29: E-tRNA
+Macromolecule #30: 16S rRNA
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20mM HEPES, 20mM MgCl2, 100mM NH4Cl, 3mM DTT |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Number grids imaged: 3 / Number real images: 12343 / Average exposure time: 2.0 sec. / Average electron dose: 1.34 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)