[English] 日本語
 Yorodumi
Yorodumi- EMDB-37413: De novo transcribing complex 10 (AdML10G), the early elongation c... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | De novo transcribing complex 10 (AdML10G), the early elongation complex with Pol II positioned 10nt downstream of TSS. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcribing complex / de novo transcription initiation / early elongation complex (EEC) / TRANSCRIPTION | |||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) / Homo sapiens (human) / synthetic construct (others) /  Amanita phalloides (death cap) Amanita phalloides (death cap) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.47 Å | |||||||||
 Authors Authors | Chen X / Liu W / Xu Y | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structural visualization of transcription initiation in action. Authors: Xizi Chen / Weida Liu / Qianmin Wang / Xinxin Wang / Yulei Ren / Xuechun Qu / Wanjun Li / Yanhui Xu /  Abstract: Transcription initiation is a complex process, and its mechanism is incompletely understood. We determined the structures of de novo transcribing complexes TC2 to TC17 with RNA polymerase II halted ...Transcription initiation is a complex process, and its mechanism is incompletely understood. We determined the structures of de novo transcribing complexes TC2 to TC17 with RNA polymerase II halted on G-less promoters when nascent RNAs reach 2 to 17 nucleotides in length, respectively. Connecting these structures generated a movie and a working model. As initially synthesized RNA grows, general transcription factors (GTFs) remain bound to the promoter and the transcription bubble expands. Nucleoside triphosphate (NTP)-driven RNA-DNA translocation and template-strand accumulation in a nearly sealed channel may promote the transition from initially transcribing complexes (ITCs) (TC2 to TC9) to early elongation complexes (EECs) (TC10 to TC17). Our study shows dynamic processes of transcription initiation and reveals why ITCs require GTFs and bubble expansion for initial RNA synthesis, whereas EECs need GTF dissociation from the promoter and bubble collapse for promoter escape. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37413.map.gz emd_37413.map.gz | 2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37413-v30.xml emd-37413-v30.xml emd-37413.xml emd-37413.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_37413.png emd_37413.png | 127.3 KB | ||
| Filedesc metadata |  emd-37413.cif.gz emd-37413.cif.gz | 4.5 KB | ||
| Others |  emd_37413_additional_1.map.gz emd_37413_additional_1.map.gz emd_37413_half_map_1.map.gz emd_37413_half_map_1.map.gz emd_37413_half_map_2.map.gz emd_37413_half_map_2.map.gz | 74.2 MB 69.9 MB 69.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37413 http://ftp.pdbj.org/pub/emdb/structures/EMD-37413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37413 | HTTPS FTP |
-Related structure data
| Related structure data |  8wakC  8walC  8wanC  8waoC  8wapC  8waqC  8warC  8wasC  8watC  8wauC  8wavC  8wawC  8waxC  8wayC  8wazC  8wb0C C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37413.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37413.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.334 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Overall map of AdML10G
| File | emd_37413_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall map of AdML10G | ||||||||||||
| Projections & Slices |
| ||||||||||||
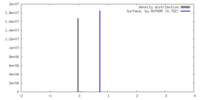
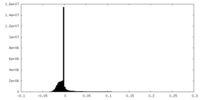
| Density Histograms |
-Half map: #1
| File | emd_37413_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_37413_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : De novo transcribing complex 10 (AdML10G), the early elongation c...
| Entire | Name: De novo transcribing complex 10 (AdML10G), the early elongation complex with Pol II positioned 10nt downstream of TSS. |
|---|---|
| Components |
|
-Supramolecule #1: De novo transcribing complex 10 (AdML10G), the early elongation c...
| Supramolecule | Name: De novo transcribing complex 10 (AdML10G), the early elongation complex with Pol II positioned 10nt downstream of TSS. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#18 |
|---|
-Supramolecule #2: RNA POLYMERASE II
| Supramolecule | Name: RNA POLYMERASE II / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #7-#18 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: TFIIF
| Supramolecule | Name: TFIIF / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: DNA
| Supramolecule | Name: DNA / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4-#5 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) / Synthetically produced: Yes |
-Supramolecule #5: RNA
| Supramolecule | Name: RNA / type: complex / ID: 5 / Parent: 1 / Macromolecule list: #6 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) / Synthetically produced: Yes |
-Supramolecule #6: Alpha-amanitin
| Supramolecule | Name: Alpha-amanitin / type: complex / ID: 6 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Amanita phalloides (death cap) Amanita phalloides (death cap) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.47 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 85241 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)