[English] 日本語
 Yorodumi
Yorodumi- EMDB-37336: Human Consensus Olfactory Receptor OR52c in apo state, OR52c only -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Consensus Olfactory Receptor OR52c in apo state, OR52c only | |||||||||
 Map data Map data | Locally refined map focused on OR52c | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Olfactory Receptor / G Protein / MEMBRANE PROTEIN / GPCR / Olfactory GPCR | |||||||||
| Function / homology | Cytochrome b562 / Cytochrome b562 / Cytochrome c/b562 / electron transport chain / periplasmic space / electron transfer activity / iron ion binding / heme binding / Soluble cytochrome b562 Function and homology information Function and homology information | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
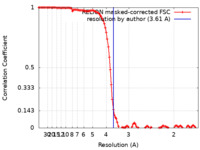
| Method | single particle reconstruction / cryo EM / Resolution: 3.61 Å | |||||||||
 Authors Authors | Choi CW / Bae J / Choi H-J / Kim J | |||||||||
| Funding support |  Korea, Republic Of, 1 items Korea, Republic Of, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Understanding the molecular mechanisms of odorant binding and activation of the human OR52 family. Authors: Chulwon Choi / Jungnam Bae / Seonghan Kim / Seho Lee / Hyunook Kang / Jinuk Kim / Injin Bang / Kiheon Kim / Won-Ki Huh / Chaok Seok / Hahnbeom Park / Wonpil Im / Hee-Jung Choi /   Abstract: Structural and mechanistic studies on human odorant receptors (ORs), key in olfactory signaling, are challenging because of their low surface expression in heterologous cells. The recent structure of ...Structural and mechanistic studies on human odorant receptors (ORs), key in olfactory signaling, are challenging because of their low surface expression in heterologous cells. The recent structure of OR51E2 bound to propionate provided molecular insight into odorant recognition, but the lack of an inactive OR structure limited understanding of the activation mechanism of ORs upon odorant binding. Here, we determined the cryo-electron microscopy structures of consensus OR52 (OR52), a representative of the OR52 family, in the ligand-free (apo) and octanoate-bound states. The apo structure of OR52 reveals a large opening between transmembrane helices (TMs) 5 and 6. A comparison between the apo and active structures of OR52 demonstrates the inward and outward movements of the extracellular and intracellular segments of TM6, respectively. These results, combined with molecular dynamics simulations and signaling assays, shed light on the molecular mechanisms of odorant binding and activation of the OR52 family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37336.map.gz emd_37336.map.gz | 62.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37336-v30.xml emd-37336-v30.xml emd-37336.xml emd-37336.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37336_fsc.xml emd_37336_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_37336.png emd_37336.png | 117.6 KB | ||
| Filedesc metadata |  emd-37336.cif.gz emd-37336.cif.gz | 6.1 KB | ||
| Others |  emd_37336_half_map_1.map.gz emd_37336_half_map_1.map.gz emd_37336_half_map_2.map.gz emd_37336_half_map_2.map.gz | 116 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37336 http://ftp.pdbj.org/pub/emdb/structures/EMD-37336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37336 | HTTPS FTP |
-Related structure data
| Related structure data |  8w77MC  8htgC  8htiC  8j46C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37336.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37336.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally refined map focused on OR52c | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.848 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map A of OR52c in apo state
| File | emd_37336_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A of OR52c in apo state | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B of OR52c in apo state
| File | emd_37336_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B of OR52c in apo state | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of Consensus Olfactory receptor OR52c and anti-bRIL Fab
| Entire | Name: Complex of Consensus Olfactory receptor OR52c and anti-bRIL Fab |
|---|---|
| Components |
|
-Supramolecule #1: Complex of Consensus Olfactory receptor OR52c and anti-bRIL Fab
| Supramolecule | Name: Complex of Consensus Olfactory receptor OR52c and anti-bRIL Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 140 KDa |
-Macromolecule #1: Human Consensus Olfactory Receptor OR52c in apo state, receptor o...
| Macromolecule | Name: Human Consensus Olfactory Receptor OR52c in apo state, receptor only,Soluble cytochrome b562 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 49.480332 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DYKDDDDAID MPTSNHTSFH PSSFLLVGIP GLESVHIWIS IPFCAMYLIA LLGNSTLLFV IKTERSLHEP MYYFLAMLAA TDLVLSTST IPKMLAIFWF NLKEISFDAC LTQMFFIHSF TGMESGVLLA MAFDRYVAIC YPLRYTTILT NKVIGKIGMA V VLRAVLLV ...String: DYKDDDDAID MPTSNHTSFH PSSFLLVGIP GLESVHIWIS IPFCAMYLIA LLGNSTLLFV IKTERSLHEP MYYFLAMLAA TDLVLSTST IPKMLAIFWF NLKEISFDAC LTQMFFIHSF TGMESGVLLA MAFDRYVAIC YPLRYTTILT NKVIGKIGMA V VLRAVLLV IPFPFLLKRL PFCGTNIIPH TYCEHMGVAK LACADIKVNI IYGLFVALLI VGLDVILIAL SYVLILRAAR RQ LADLEDN WETLNDNLKV IEKADNAAQV KDALTKMRAA ALDAQKATPP KLEDKSPDSP EMKDFRHGFD ILVGQIDDAL KLA NEGKVK EAQAAAEQLK TTRNAYIQKY LERARSTLLK ALSTCGSHIC VILAFYTPAF FSFLTHRFGH HIPPYIHILL ANLY LLVPP MLNPIIYGVK TKQIRERVLK IFFKKKASGL EVLFQ UniProtKB: Soluble cytochrome b562 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV / Details: blot time 3 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Average electron dose: 68.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8w77: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)