+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human TMEM87A A308M | |||||||||
 Map data Map data | cryo-EM map of human TMEM87A with A308M mutation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | non-selective cation channel / ion channel / membrane protein / Golgi-localized protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationdetection of mechanical stimulus involved in sensory perception of touch / retrograde transport, endosome to Golgi / Golgi cisterna membrane / RHOA GTPase cycle / ruffle / bioluminescence / generation of precursor metabolites and energy / cellular response to mechanical stimulus / Golgi membrane / Golgi apparatus ...detection of mechanical stimulus involved in sensory perception of touch / retrograde transport, endosome to Golgi / Golgi cisterna membrane / RHOA GTPase cycle / ruffle / bioluminescence / generation of precursor metabolites and energy / cellular response to mechanical stimulus / Golgi membrane / Golgi apparatus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human adenovirus C serotype 2 Human adenovirus C serotype 2 | |||||||||
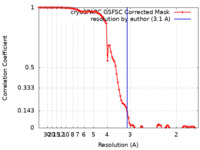
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Han A / Kim HM | |||||||||
| Funding support |  Korea, Republic Of, 1 items Korea, Republic Of, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of human TMEM87A, PE-bound Authors: Han A / Kim HM | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37069.map.gz emd_37069.map.gz | 41.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37069-v30.xml emd-37069-v30.xml emd-37069.xml emd-37069.xml | 20.6 KB 20.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37069_fsc.xml emd_37069_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_37069.png emd_37069.png | 33.4 KB | ||
| Filedesc metadata |  emd-37069.cif.gz emd-37069.cif.gz | 7.3 KB | ||
| Others |  emd_37069_half_map_1.map.gz emd_37069_half_map_1.map.gz emd_37069_half_map_2.map.gz emd_37069_half_map_2.map.gz | 77.8 MB 77.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37069 http://ftp.pdbj.org/pub/emdb/structures/EMD-37069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37069 | HTTPS FTP |
-Validation report
| Summary document |  emd_37069_validation.pdf.gz emd_37069_validation.pdf.gz | 848.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_37069_full_validation.pdf.gz emd_37069_full_validation.pdf.gz | 848.2 KB | Display | |
| Data in XML |  emd_37069_validation.xml.gz emd_37069_validation.xml.gz | 17.4 KB | Display | |
| Data in CIF |  emd_37069_validation.cif.gz emd_37069_validation.cif.gz | 22.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37069 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37069 | HTTPS FTP |
-Related structure data
| Related structure data |  8kb4MC  8hsiC  8httC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37069.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37069.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM map of human TMEM87A with A308M mutation | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.848 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: cryo-EM half A map of human TMEM87A with A308M mutation
| File | emd_37069_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half A map of human TMEM87A with A308M mutation | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cryo-EM half B map of human TMEM87A with A308M mutation
| File | emd_37069_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half_B_map of human TMEM87A with A308M mutation | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Transmembrane protein 87A A308M with GFP tag and Twin-strep tag
| Entire | Name: Transmembrane protein 87A A308M with GFP tag and Twin-strep tag |
|---|---|
| Components |
|
-Supramolecule #1: Transmembrane protein 87A A308M with GFP tag and Twin-strep tag
| Supramolecule | Name: Transmembrane protein 87A A308M with GFP tag and Twin-strep tag type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: Transmembrane protein 87A A308M with GFP tag and Twin-strep tag purified with detergent LMNG/CHS |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Transmembrane protein 87A,EGFP
| Macromolecule | Name: Transmembrane protein 87A,EGFP / type: protein_or_peptide / ID: 1 Details: Transmembrane protein 87A A308M with GFP and twin strep tag Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human adenovirus C serotype 2 Human adenovirus C serotype 2 |
| Molecular weight | Theoretical: 96.983539 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAAAAWLQVL PVILLLLGAH PSPLSFFSAG PATVAAADRS KWHIPIPSGK NYFSFGKILF RNTTIFLKFD GEPCDLSLNI TWYLKSADC YNEIYNFKAE EVELYLEKLK EKRGLSGKYQ TSSKLFQNCS ELFKTQTFSG DFMHRLPLLG EKQEAKENGT N LTFIGDKT ...String: MAAAAWLQVL PVILLLLGAH PSPLSFFSAG PATVAAADRS KWHIPIPSGK NYFSFGKILF RNTTIFLKFD GEPCDLSLNI TWYLKSADC YNEIYNFKAE EVELYLEKLK EKRGLSGKYQ TSSKLFQNCS ELFKTQTFSG DFMHRLPLLG EKQEAKENGT N LTFIGDKT AMHEPLQTWQ DAPYIFIVHI GISSSKESSK ENSLSNLFTM TVEVKGPYEY LTLEDYPLMI FFMVMCIVYV LF GVLWLAW SACYWRDLLR IQFWIGAVIF LGMLEKAVFY AEFQNIRYKG ESVQGALILA ELLSAVKRSL MRTLVIIVSL GYG IVKPRL GVTLHKVVVA GALYLLFSGM EGVLRVTGAQ TDLASLAFIP LAFLDTALCW WIFISLTQTM KLLKLRRNIV KLSL YRHFT NTLILAVAAS IVFIIWTTMK FRIVTCQSDW RELWVDDAIW RLLFSMILFV IMVLWRPSAN NQRFAFSPLS EEEEE DEQK EPMLKESFEG MKMRSTKQEP NGNSKVNKAQ EDDLKWVEEN VPSSVTDVAL PALLDSDEER MITHFERSKM ELKENL YFQ GNPAFLYKVV DPVVSKGEEL FTGVVPILVE LDGDVNGHKF SVSGEGEGDA TYGKLTLKFI CTTGKLPVPW PTLVTTL TY GVQCFSRYPD HMKQHDFFKS AMPEGYVQER TIFFKDDGNY KTRAEVKFEG DTLVNRIELK GIDFKEDGNI LGHKLEYN Y NSHNVYIMAD KQKNGIKVNF KIRHNIEDGS VQLADHYQQN TPIGDGPVLL PDNHYLSTQS ALSKDPNEKR DHMVLLEFV TAAGITLGMD ELYKEFLVPR GSSRSAWSHP QFEKGGGSGG GSGGSAWSHP QFEK UniProtKB: Transmembrane protein 87A, EGFP |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: (9R,12R)-15-amino-12-hydroxy-6,12-dioxo-7,11,13-trioxa-12lambda~5...
| Macromolecule | Name: (9R,12R)-15-amino-12-hydroxy-6,12-dioxo-7,11,13-trioxa-12lambda~5~-phosphapentadecan-9-yl undecanoate type: ligand / ID: 3 / Number of copies: 1 / Formula: 65I |
|---|---|
| Molecular weight | Theoretical: 481.56 Da |
| Chemical component information | 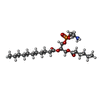 ChemComp-65I: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 9 Component:
Details: 50mM HEPES pH 7.5, 250mM NaCl, 0.01% (w/v) LMNG, 0.002% (w/v) CHS | |||||||||||||||
| Grid | Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.015 kPa Details: The grid was negatively glow-discharged for 60 sec with 15mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Coma free - Residual tilt: 10.0 mrad |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Details | Using Zemlin tableau in sherpa |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 4565 / Average exposure time: 6.09 sec. / Average electron dose: 67.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 58900 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Chimera Fit in map tool was used for initial local fitting. Then, Real-space refinement with the rigid body option in PHENIX was used for flexible fitting. |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 40 |
| Output model |  PDB-8kb4: |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X