[English] 日本語
 Yorodumi
Yorodumi- EMDB-3655: Cryo-Electron Microscopy Structure of the Macrobrachium rosenberg... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3655 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-Electron Microscopy Structure of the Macrobrachium rosenbergii Nodavirus Capsid at 7 Angstroms Resolution | |||||||||
 Map data Map data | Macrobrachium rosenbergii nodavirus virus-like particle. | |||||||||
 Sample Sample | Macrobrachium rosenbergii Nodavirus Capsid != Macrobrachium rosenbergii Nodavirus Macrobrachium rosenbergii Nodavirus Capsid
| |||||||||
| Biological species |  Macrobrachium rosenbergii Nodavirus Macrobrachium rosenbergii Nodavirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Bhella D / Ho KL | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2017 Journal: Sci Rep / Year: 2017Title: Cryo-Electron Microscopy Structure of the Macrobrachium rosenbergii Nodavirus Capsid at 7 Angstroms Resolution. Authors: Kok Lian Ho / Chare Li Kueh / Poay Ling Beh / Wen Siang Tan / David Bhella /   Abstract: White tail disease in the giant freshwater prawn Macrobrachium rosenbergii causes significant economic losses in shrimp farms and hatcheries and poses a threat to food-security in many developing ...White tail disease in the giant freshwater prawn Macrobrachium rosenbergii causes significant economic losses in shrimp farms and hatcheries and poses a threat to food-security in many developing countries. Outbreaks of Macrobrachium rosenbergii nodavirus (MrNV), the causative agent of white tail disease (WTD) are associated with up to 100% mortality rates. There are no interventions available to treat or prevent MrNV disease however. Here we show the structure of MrNV virus-like particles (VLPs) produced by recombinant expression of the capsid protein, using cryogenic electron microscopy. Our data show that MrNV VLPs package nucleic acids in a manner reminiscent of other known nodavirus structures. The structure of the capsid however shows striking differences from insect and fish infecting nodaviruses, which have been shown to assemble trimer-clustered T = 3 icosahedral virus particles. MrNV particles have pronounced dimeric blade-shaped spikes extending up to 6 nm from the outer surface of the capsid shell. Our structural analysis supports the assertion that MrNV may belong to a new genus of the Nodaviridae. Moreover, our study provides the first structural view of an important pathogen affecting aquaculture industries across the world. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3655.map.gz emd_3655.map.gz | 66.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3655-v30.xml emd-3655-v30.xml emd-3655.xml emd-3655.xml | 11.4 KB 11.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3655_fsc.xml emd_3655_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_3655.png emd_3655.png | 198.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3655 http://ftp.pdbj.org/pub/emdb/structures/EMD-3655 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3655 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3655 | HTTPS FTP |
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3655.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3655.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Macrobrachium rosenbergii nodavirus virus-like particle. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
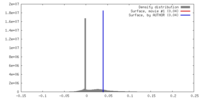
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Macrobrachium rosenbergii Nodavirus Capsid
| Entire | Name: Macrobrachium rosenbergii Nodavirus Capsid |
|---|---|
| Components |
|
-Supramolecule #1: Macrobrachium rosenbergii Nodavirus
| Supramolecule | Name: Macrobrachium rosenbergii Nodavirus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 222557 / Sci species name: Macrobrachium rosenbergii Nodavirus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Macrobrachium rosenbergii (giant freshwater prawn) Macrobrachium rosenbergii (giant freshwater prawn) |
| Virus shell | Shell ID: 1 / Diameter: 400.0 Å / T number (triangulation number): 3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Grid | Model: Protochips C-flat R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR Details: C-flats had an additional carbon evaporation applied |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Blot for 4 seconds, Blot force 25. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FSC |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-20 (5k x 3k) / Detector mode: INTEGRATING / Digitization - Frames/image: 2-60 / Number grids imaged: 4 / Number real images: 281 / Average exposure time: 3.0 sec. / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)