[English] 日本語
 Yorodumi
Yorodumi- EMDB-3565: Cowpea mosaic virus top component (CPMV-T) - naturally occurring ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3565 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cowpea mosaic virus top component (CPMV-T) - naturally occurring empty particles | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CPMV-M / plant virus / picornavirales / virus | |||||||||
| Function / homology |  Function and homology information Function and homology informationtransport of virus in host, cell to cell / host cell plasmodesma / T=3 icosahedral viral capsid / symbiont-mediated suppression of host innate immune response / GTP binding / host cell nucleus / structural molecule activity / DNA binding / RNA binding Similarity search - Function | |||||||||
| Biological species |   Cowpea mosaic virus Cowpea mosaic virus | |||||||||
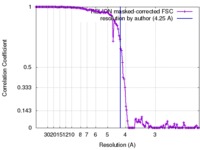
| Method | single particle reconstruction / cryo EM / Resolution: 4.25 Å | |||||||||
 Authors Authors | Hesketh EL / Meshcheriakova Y | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2017 Journal: Sci Rep / Year: 2017Title: The structures of a naturally empty cowpea mosaic virus particle and its genome-containing counterpart by cryo-electron microscopy. Authors: Emma L Hesketh / Yulia Meshcheriakova / Rebecca F Thompson / George P Lomonossoff / Neil A Ranson /  Abstract: Cowpea mosaic virus (CPMV) is a picorna-like plant virus. As well as an intrinsic interest in CPMV as a plant pathogen, CPMV is of major interest in biotechnology applications such as nanotechnology. ...Cowpea mosaic virus (CPMV) is a picorna-like plant virus. As well as an intrinsic interest in CPMV as a plant pathogen, CPMV is of major interest in biotechnology applications such as nanotechnology. Here, we report high resolution cryo electron microscopy (cryo-EM) maps of wild type CPMV containing RNA-2, and of naturally-formed empty CPMV capsids. The resolution of these structures is sufficient to visualise large amino acids. We have refined an atomic model for each map and identified an essential amino acid involved in genome encapsidation. This work has furthered our knowledge of Picornavirales genome encapsidation and will assist further work in the development of CPMV as a biotechnological tool. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3565.map.gz emd_3565.map.gz | 45.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3565-v30.xml emd-3565-v30.xml emd-3565.xml emd-3565.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3565_fsc.xml emd_3565_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_3565.png emd_3565.png | 267.2 KB | ||
| Filedesc metadata |  emd-3565.cif.gz emd-3565.cif.gz | 5.8 KB | ||
| Others |  emd_3565_additional.map.gz emd_3565_additional.map.gz | 165.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3565 http://ftp.pdbj.org/pub/emdb/structures/EMD-3565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3565 | HTTPS FTP |
-Related structure data
| Related structure data |  5mshMC  3562C  8609C  5ms1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3565.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3565.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 3565 additional.map
| File | emd_3565_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cowpea mosaic virus
| Entire | Name:   Cowpea mosaic virus Cowpea mosaic virus |
|---|---|
| Components |
|
-Supramolecule #1: Cowpea mosaic virus
| Supramolecule | Name: Cowpea mosaic virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 12264 / Sci species name: Cowpea mosaic virus / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Vigna unguiculata (cowpea) Vigna unguiculata (cowpea) |
-Macromolecule #1: Cowpea mosaic virus small subunit
| Macromolecule | Name: Cowpea mosaic virus small subunit / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Cowpea mosaic virus / Strain: SB Cowpea mosaic virus / Strain: SB |
| Molecular weight | Theoretical: 20.961564 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPVCAEASDV YSPCMIASTP PAPFSDVTAV TFDLINGKIT PVGDDNWNTH IYNPPIMNVL RTAAWKSGTI HVQLNVRGAG VKRADWDGQ VFVYLRQSMN PESYDARTFV ISQPGSAMLN FSFDIIGPNS GFEFAESPWA NQTTWYLECV ATNPRQIQQF E VNMRFDPN FRVAGNILMP PFPLSTETPP L UniProtKB: RNA2 polyprotein |
-Macromolecule #2: Cowpea mosaic virus large subunit
| Macromolecule | Name: Cowpea mosaic virus large subunit / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Cowpea mosaic virus / Strain: SB Cowpea mosaic virus / Strain: SB |
| Molecular weight | Theoretical: 40.858434 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEQNLFALSL DDTSSVRGSL LDTKFAQTRV LLSKAMAGGD VLLDEYLYDV VNGQDFRATV AFLRTHVITG KIKVTATTNI SDNSGCCLM LAINSGVRGK YSTDVYTICS QDSMTWNPGC KKNFSFTFNP NPCGDSWSAE MISRSRVRMT VICVSGWTLS P TTDVIAKL ...String: MEQNLFALSL DDTSSVRGSL LDTKFAQTRV LLSKAMAGGD VLLDEYLYDV VNGQDFRATV AFLRTHVITG KIKVTATTNI SDNSGCCLM LAINSGVRGK YSTDVYTICS QDSMTWNPGC KKNFSFTFNP NPCGDSWSAE MISRSRVRMT VICVSGWTLS P TTDVIAKL DWSIVNEKCE PTIYHLADCQ NWLPLNRWMG KLTFPQGVTS EVRRMPLSIG GGAGATQAFL ANMPNSWISM WR YFRGELH FEVTKMSSPY IKATVTFLIA FGNLSDAFGF YESFPHRIVQ FAEVEEKCTL VFSQQEFVTA WSTQVNPRTT LEA DGCPYL YAIIHDSTTG TISGDFNLGV KLVGIKDFCG IGSNPGIDGS RLL UniProtKB: RNA2 polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.9 mg/mL |
|---|---|
| Buffer | pH: 7.8 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Number grids imaged: 1 / Number real images: 1759 / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 10.0 µm / Nominal defocus min: 0.35000000000000003 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-5msh: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)