[English] 日本語
 Yorodumi
Yorodumi- EMDB-35620: The Rubisco assembly intermidiate of Rubisco large subunit (RbcL)... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Rubisco assembly intermidiate of Rubisco large subunit (RbcL) and Arabidopsis thaliana Rubisco accumulation factor 1 (AtRaf1) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Rubisco assembly intermidiate / CHAPERONE / LYASE-CHAPERONE complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationribulose bisphosphate carboxylase complex assembly / photorespiration / carboxysome / ribulose-bisphosphate carboxylase / ribulose-bisphosphate carboxylase activity / reductive pentose-phosphate cycle / chloroplast stroma / chloroplast / monooxygenase activity / magnesium ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |   Synechococcus elongatus PCC 6301 (bacteria) Synechococcus elongatus PCC 6301 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Wang R / Song H / Zhang W / Wang N / Zhang S / Shao R | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Plant / Year: 2023 Journal: Mol Plant / Year: 2023Title: Structural insights into the functions of Raf1 and Bsd2 in hexadecameric Rubisco assembly. Authors: Ran Wang / Hui Song / Wenjuan Zhang / Ning Wang / Shijia Zhang / Ruiqi Shao / Cuimin Liu /  Abstract: Hexadecameric form I Rubisco, which consisting consists of eight large (RbcL) and eight small (RbcS) subunits, is the most abundant enzyme on earth. Extensive efforts to engineer an improved Rubisco ...Hexadecameric form I Rubisco, which consisting consists of eight large (RbcL) and eight small (RbcS) subunits, is the most abundant enzyme on earth. Extensive efforts to engineer an improved Rubisco to speed up its catalytic efficiency and ultimately increase agricultural productivity. However, difficulties with correct folding and assembly in foreign hosts or in vitro have hampered the genetic manipulation of hexadecameric Rubisco. In this study, we reconstituted Synechococcus sp. PCC6301 Rubisco in vitro using the chaperonin system and assembly factors from cyanobacteria and Arabidopsis thaliana (At). Rubisco holoenzyme was produced in the presence of cyanobacterial Rubisco accumulation factor 1 (Raf1) alone or both AtRaf1 and bundle-sheath defective-2 (AtBsd2) from Arabidopsis. RbcL released from GroEL is assembly capable in the presence of ATP, and AtBsd2 functions downstream of AtRaf1. Cryo-EM structures of RbcL-AtRaf1, RbcL-AtRaf1-AtBsd2, and RbcL revealed that the interactions between RbcL and AtRaf1 are looser than those between prokaryotic RbcL and Raf1, with AtRaf1 tilting 7° farther away from RbcL. AtBsd2 stabilizes the flexible regions of RbcL, including the N and C termini, the 60s loop, and loop 6. Using these data, combined with previous findings, we propose the possible biogenesis pathways of prokaryotic and eukaryotic Rubisco. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35620.map.gz emd_35620.map.gz | 41.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35620-v30.xml emd-35620-v30.xml emd-35620.xml emd-35620.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35620.png emd_35620.png | 51.6 KB | ||
| Filedesc metadata |  emd-35620.cif.gz emd-35620.cif.gz | 5.8 KB | ||
| Others |  emd_35620_half_map_1.map.gz emd_35620_half_map_1.map.gz emd_35620_half_map_2.map.gz emd_35620_half_map_2.map.gz | 218.5 MB 218.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35620 http://ftp.pdbj.org/pub/emdb/structures/EMD-35620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35620 | HTTPS FTP |
-Related structure data
| Related structure data |  8iojMC  8ilbC  8ilmC  8io2C  8iolC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35620.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35620.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.52 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35620_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
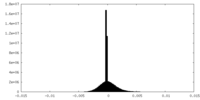
| Density Histograms |
-Half map: #1
| File | emd_35620_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : the Rubisco assembly intermediate of Rubisco large subunit (RbcL)...
| Entire | Name: the Rubisco assembly intermediate of Rubisco large subunit (RbcL) and Arabidopsis thaliana Rubisco accumulation factor 1 (AtRaf1) |
|---|---|
| Components |
|
-Supramolecule #1: the Rubisco assembly intermediate of Rubisco large subunit (RbcL)...
| Supramolecule | Name: the Rubisco assembly intermediate of Rubisco large subunit (RbcL) and Arabidopsis thaliana Rubisco accumulation factor 1 (AtRaf1) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ribulose bisphosphate carboxylase large chain
| Macromolecule | Name: Ribulose bisphosphate carboxylase large chain / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:  Synechococcus elongatus PCC 6301 (bacteria) / Strain: ATCC 27144 / PCC 6301 / SAUG 1402/1 Synechococcus elongatus PCC 6301 (bacteria) / Strain: ATCC 27144 / PCC 6301 / SAUG 1402/1 |
| Molecular weight | Theoretical: 52.516605 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPKTQSAAGY KAGVKDYKLT YYTPDYTPKD TDLLAAFRFS PQPGVPADEA GAAIAAESST GTWTTVWTDL LTDMDRYKGK CYHIEPVQG EENSYFAFIA YPLDLFEEGS VTNILTSIVG NVFGFKAIRS LRLEDIRFPV ALVKTFQGPP HGIQVERDLL N KYGRPMLG ...String: MPKTQSAAGY KAGVKDYKLT YYTPDYTPKD TDLLAAFRFS PQPGVPADEA GAAIAAESST GTWTTVWTDL LTDMDRYKGK CYHIEPVQG EENSYFAFIA YPLDLFEEGS VTNILTSIVG NVFGFKAIRS LRLEDIRFPV ALVKTFQGPP HGIQVERDLL N KYGRPMLG CTIKPKLGLS AKNYGRAVYE CLRGGLDFTK DDENINSQPF QRWRDRFLFV ADAIHKSQAE TGEIKGHYLN VT APTCEEM MKRAEFAKEL GMPIIMHDFL TAGFTANTTL AKWCRDNGVL LHIHRAMHAV IDRQRNHGIH FRVLAKCLRL SGG DHLHSG TVVGKLEGDK ASTLGFVDLM REDHIEADRS RGVFFTQDWA SMPGVLPVAS GGIHVWHMPA LVEIFGDDSV LQFG GGTLG HPWGNAPGAT ANRVALEACV QARNEGRDLY REGGDILREA GKWSPELAAA LDLWKEIKFE FETMDKL UniProtKB: Ribulose bisphosphate carboxylase large chain |
-Macromolecule #2: Rubisco accumulation factor 1.2, chloroplastic
| Macromolecule | Name: Rubisco accumulation factor 1.2, chloroplastic / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.823639 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QQLYQPFRPP SSPIPTQFRS LDSAGKIEIL AGRMALWFEY APLISSLYTD GFTPPTIEEL TGISSIEQNR LIVGAQVRDS ILQSIHEPE LISAFDTGGA ELLYEIRLLS TTQRVAAATF IIDRNIDSKG AQDLARAIKD YPNRRGDVGW LDFDYNLPGD C LSFLYYRQ ...String: QQLYQPFRPP SSPIPTQFRS LDSAGKIEIL AGRMALWFEY APLISSLYTD GFTPPTIEEL TGISSIEQNR LIVGAQVRDS ILQSIHEPE LISAFDTGGA ELLYEIRLLS TTQRVAAATF IIDRNIDSKG AQDLARAIKD YPNRRGDVGW LDFDYNLPGD C LSFLYYRQ SRENKNPSDQ RTSMLLQALG VAESEKAKNR LNTEL UniProtKB: Rubisco accumulation factor 1.2, chloroplastic |
-Macromolecule #3: Rubisco accumulation factor 1.2, chloroplastic
| Macromolecule | Name: Rubisco accumulation factor 1.2, chloroplastic / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.639338 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: RIPVVRLKFG EVAEATSVVV LPVCKAEEGE KKILEAPMEI IAGGDFKVVE AEKGWKRWVV LPSWNPVAAI GKGGVAVSFR DDRKVLPWD GKEEPLLVVA DRVRNVVEAD DGYYLVVAEN GLKLEKGSDL KAREVKESLG MVVLVVRPPR ED UniProtKB: Rubisco accumulation factor 1.2, chloroplastic |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 239291 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)