[English] 日本語
 Yorodumi
Yorodumi- EMDB-35605: The Rubisco assembly intermidate of Arabidopsis thaliana Rubisco ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Rubisco assembly intermidate of Arabidopsis thaliana Rubisco accumulation factor 1 (AtRaf1) and Rubisco large subunit (RbcL) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Rubisco assembly intermediate / CHAPERONE / LYASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationribulose bisphosphate carboxylase complex assembly / photorespiration / carboxysome / ribulose-bisphosphate carboxylase / ribulose-bisphosphate carboxylase activity / reductive pentose-phosphate cycle / chloroplast stroma / chloroplast / monooxygenase activity / magnesium ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |   Synechococcus sp. (strain ATCC 27144 / PCC 6301 / SAUG 1402/1) (bacteria) Synechococcus sp. (strain ATCC 27144 / PCC 6301 / SAUG 1402/1) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Wang R / Song H / Zhang W / Wang N / Zhang S / Shao R | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Plant / Year: 2023 Journal: Mol Plant / Year: 2023Title: Structural insights into the functions of Raf1 and Bsd2 in hexadecameric Rubisco assembly. Authors: Ran Wang / Hui Song / Wenjuan Zhang / Ning Wang / Shijia Zhang / Ruiqi Shao / Cuimin Liu /  Abstract: Hexadecameric form I Rubisco, which consisting consists of eight large (RbcL) and eight small (RbcS) subunits, is the most abundant enzyme on earth. Extensive efforts to engineer an improved Rubisco ...Hexadecameric form I Rubisco, which consisting consists of eight large (RbcL) and eight small (RbcS) subunits, is the most abundant enzyme on earth. Extensive efforts to engineer an improved Rubisco to speed up its catalytic efficiency and ultimately increase agricultural productivity. However, difficulties with correct folding and assembly in foreign hosts or in vitro have hampered the genetic manipulation of hexadecameric Rubisco. In this study, we reconstituted Synechococcus sp. PCC6301 Rubisco in vitro using the chaperonin system and assembly factors from cyanobacteria and Arabidopsis thaliana (At). Rubisco holoenzyme was produced in the presence of cyanobacterial Rubisco accumulation factor 1 (Raf1) alone or both AtRaf1 and bundle-sheath defective-2 (AtBsd2) from Arabidopsis. RbcL released from GroEL is assembly capable in the presence of ATP, and AtBsd2 functions downstream of AtRaf1. Cryo-EM structures of RbcL-AtRaf1, RbcL-AtRaf1-AtBsd2, and RbcL revealed that the interactions between RbcL and AtRaf1 are looser than those between prokaryotic RbcL and Raf1, with AtRaf1 tilting 7° farther away from RbcL. AtBsd2 stabilizes the flexible regions of RbcL, including the N and C termini, the 60s loop, and loop 6. Using these data, combined with previous findings, we propose the possible biogenesis pathways of prokaryotic and eukaryotic Rubisco. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35605.map.gz emd_35605.map.gz | 46.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35605-v30.xml emd-35605-v30.xml emd-35605.xml emd-35605.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35605.png emd_35605.png | 39.9 KB | ||
| Filedesc metadata |  emd-35605.cif.gz emd-35605.cif.gz | 5.8 KB | ||
| Others |  emd_35605_half_map_1.map.gz emd_35605_half_map_1.map.gz emd_35605_half_map_2.map.gz emd_35605_half_map_2.map.gz | 212.7 MB 212.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35605 http://ftp.pdbj.org/pub/emdb/structures/EMD-35605 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35605 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35605 | HTTPS FTP |
-Related structure data
| Related structure data |  8io2MC  8ilbC  8ilmC  8iojC  8iolC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35605.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35605.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.52 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35605_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
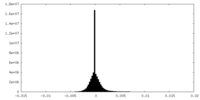
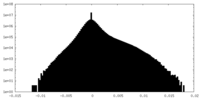
| Density Histograms |
-Half map: #1
| File | emd_35605_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : the complex of RbcL and Raf1 (L8F8)
| Entire | Name: the complex of RbcL and Raf1 (L8F8) |
|---|---|
| Components |
|
-Supramolecule #1: the complex of RbcL and Raf1 (L8F8)
| Supramolecule | Name: the complex of RbcL and Raf1 (L8F8) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ribulose bisphosphate carboxylase large chain
| Macromolecule | Name: Ribulose bisphosphate carboxylase large chain / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:  Synechococcus sp. (strain ATCC 27144 / PCC 6301 / SAUG 1402/1) (bacteria) Synechococcus sp. (strain ATCC 27144 / PCC 6301 / SAUG 1402/1) (bacteria)Strain: ATCC 27144 / PCC 6301 / SAUG 1402/1 |
| Molecular weight | Theoretical: 52.38541 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PKTQSAAGYK AGVKDYKLTY YTPDYTPKDT DLLAAFRFSP QPGVPADEAG AAIAAESSTG TWTTVWTDLL TDMDRYKGKC YHIEPVQGE ENSYFAFIAY PLDLFEEGSV TNILTSIVGN VFGFKAIRSL RLEDIRFPVA LVKTFQGPPH GIQVERDLLN K YGRPMLGC ...String: PKTQSAAGYK AGVKDYKLTY YTPDYTPKDT DLLAAFRFSP QPGVPADEAG AAIAAESSTG TWTTVWTDLL TDMDRYKGKC YHIEPVQGE ENSYFAFIAY PLDLFEEGSV TNILTSIVGN VFGFKAIRSL RLEDIRFPVA LVKTFQGPPH GIQVERDLLN K YGRPMLGC TIKPKLGLSA KNYGRAVYEC LRGGLDFTKD DENINSQPFQ RWRDRFLFVA DAIHKSQAET GEIKGHYLNV TA PTCEEMM KRAEFAKELG MPIIMHDFLT AGFTANTTLA KWCRDNGVLL HIHRAMHAVI DRQRNHGIHF RVLAKCLRLS GGD HLHSGT VVGKLEGDKA STLGFVDLMR EDHIEADRSR GVFFTQDWAS MPGVLPVASG GIHVWHMPAL VEIFGDDSVL QFGG GTLGH PWGNAPGATA NRVALEACVQ ARNEGRDLYR EGGDILREAG KWSPELAAAL DLWKEIKFEF ETMDKL UniProtKB: Ribulose bisphosphate carboxylase large chain |
-Macromolecule #2: Rubisco accumulation factor 1.2, chloroplastic
| Macromolecule | Name: Rubisco accumulation factor 1.2, chloroplastic / type: protein_or_peptide / ID: 2 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.43673 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SPIPTQFRSL DSAGKIEILA GRMALWFEYA PLISSLYTDG FTPPTIEELT GISSIEQNRL IVGAQVRDSI LQSIHEPELI SAFDTGGAE LLYEIRLLST TQRVAAATFI IDRNIDSKGA QDLARAIKDY PNRRGDVGWL DFDYNLPGDC LSFLYYRQSR E NKNPSDQR ...String: SPIPTQFRSL DSAGKIEILA GRMALWFEYA PLISSLYTDG FTPPTIEELT GISSIEQNRL IVGAQVRDSI LQSIHEPELI SAFDTGGAE LLYEIRLLST TQRVAAATFI IDRNIDSKGA QDLARAIKDY PNRRGDVGWL DFDYNLPGDC LSFLYYRQSR E NKNPSDQR TSMLLQALGV AESEKAKNRL NTELYGDRIP VVRLKFGEVA EATSVVVLPV CKAEEGEKKI LEAPMEIIAG GD FKVVEAE KGWKRWVVLP SWNPVAAIGK GGVAVSFRDD RKVLPWDGKE EPLLVVADRV RNVVEADDGY YLVVAENGLK LEK GSDLKA REVKESLGMV VLVVRPPRED UniProtKB: Rubisco accumulation factor 1.2, chloroplastic |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 535498 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)