+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of the Rpd3L complex in Ume1 part | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Guo Z / Zhan X / Wang C | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structure of a SIN3-HDAC complex from budding yeast. Authors: Zhouyan Guo / Chen Chu / Yichen Lu / Xiaofeng Zhang / Yihang Xiao / Mingxuan Wu / Shuaixin Gao / Catherine C L Wong / Xiechao Zhan / Chengcheng Wang /   Abstract: SIN3-HDAC (histone deacetylases) complexes have important roles in facilitating local histone deacetylation to regulate chromatin accessibility and gene expression. Here, we present the cryo-EM ...SIN3-HDAC (histone deacetylases) complexes have important roles in facilitating local histone deacetylation to regulate chromatin accessibility and gene expression. Here, we present the cryo-EM structure of the budding yeast SIN3-HDAC complex Rpd3L at an average resolution of 2.6 Å. The structure reveals that two distinct arms (ARM1 and ARM2) hang on a T-shaped scaffold formed by two coiled-coil domains. In each arm, Sin3 interacts with different subunits to create a different environment for the histone deacetylase Rpd3. ARM1 is in the inhibited state with the active site of Rpd3 blocked, whereas ARM2 is in an open conformation with the active site of Rpd3 exposed to the exterior space. The observed asymmetric architecture of Rpd3L is different from those of available structures of other class I HDAC complexes. Our study reveals the organization mechanism of the SIN3-HDAC complex and provides insights into the interaction pattern by which it targets histone deacetylase to chromatin. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34939.map.gz emd_34939.map.gz | 218.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34939-v30.xml emd-34939-v30.xml emd-34939.xml emd-34939.xml | 12.9 KB 12.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34939.png emd_34939.png | 41.1 KB | ||
| Others |  emd_34939_half_map_1.map.gz emd_34939_half_map_1.map.gz emd_34939_half_map_2.map.gz emd_34939_half_map_2.map.gz | 192 MB 191.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34939 http://ftp.pdbj.org/pub/emdb/structures/EMD-34939 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34939 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34939 | HTTPS FTP |
-Validation report
| Summary document |  emd_34939_validation.pdf.gz emd_34939_validation.pdf.gz | 559.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34939_full_validation.pdf.gz emd_34939_full_validation.pdf.gz | 558.6 KB | Display | |
| Data in XML |  emd_34939_validation.xml.gz emd_34939_validation.xml.gz | 15.6 KB | Display | |
| Data in CIF |  emd_34939_validation.cif.gz emd_34939_validation.cif.gz | 18.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34939 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34939 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34939 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34939 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34939.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34939.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||
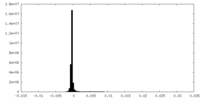
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_34939_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
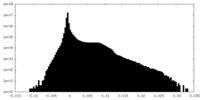
| Density Histograms |
-Half map: #2
| File | emd_34939_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
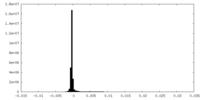
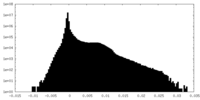
| Density Histograms |
- Sample components
Sample components
-Entire : The Rpd3L complex
| Entire | Name: The Rpd3L complex |
|---|---|
| Components |
|
-Supramolecule #1: The Rpd3L complex
| Supramolecule | Name: The Rpd3L complex / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#11 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 455965 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)