[English] 日本語
 Yorodumi
Yorodumi- EMDB-34816: Cryo-EM structure of Mycobacterium tuberculosis transcription ini... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Mycobacterium tuberculosis transcription initiation complex with transcription factor GlnR | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription factor / TRANSCRIPTION-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to water / Antimicrobial action and antimicrobial resistance in Mtb / sigma factor antagonist complex / phosphorelay response regulator activity / sigma factor activity / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / nitrate assimilation / peptidoglycan-based cell wall / DNA-directed RNA polymerase complex ...response to water / Antimicrobial action and antimicrobial resistance in Mtb / sigma factor antagonist complex / phosphorelay response regulator activity / sigma factor activity / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / nitrate assimilation / peptidoglycan-based cell wall / DNA-directed RNA polymerase complex / DNA-templated transcription initiation / protein-DNA complex / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / protein dimerization activity / transcription cis-regulatory region binding / response to antibiotic / regulation of DNA-templated transcription / DNA-templated transcription / positive regulation of DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) | |||||||||
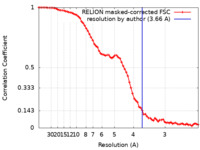
| Method | single particle reconstruction / cryo EM / Resolution: 3.66 Å | |||||||||
 Authors Authors | Lin W / Shi J / Xu JC | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structural insights into the transcription activation mechanism of the global regulator GlnR from actinobacteria. Authors: Jing Shi / Zhenzhen Feng / Juncao Xu / Fangfang Li / Yuqiong Zhang / Aijia Wen / Fulin Wang / Qian Song / Lu Wang / Hong Cui / Shujuan Tong / Peiying Chen / Yejin Zhu / Guoping Zhao / Shuang ...Authors: Jing Shi / Zhenzhen Feng / Juncao Xu / Fangfang Li / Yuqiong Zhang / Aijia Wen / Fulin Wang / Qian Song / Lu Wang / Hong Cui / Shujuan Tong / Peiying Chen / Yejin Zhu / Guoping Zhao / Shuang Wang / Yu Feng / Wei Lin /  Abstract: In actinobacteria, an OmpR/PhoB subfamily protein called GlnR acts as an orphan response regulator and globally coordinates the expression of genes responsible for nitrogen, carbon, and phosphate ...In actinobacteria, an OmpR/PhoB subfamily protein called GlnR acts as an orphan response regulator and globally coordinates the expression of genes responsible for nitrogen, carbon, and phosphate metabolism in actinobacteria. Although many researchers have attempted to elucidate the mechanisms of GlnR-dependent transcription activation, progress is impeded by lacking of an overall structure of GlnR-dependent transcription activation complex (GlnR-TAC). Here, we report a co-crystal structure of the C-terminal DNA-binding domain of GlnR (GlnR_DBD) in complex with its regulatory -element DNA and a cryo-EM structure of GlnR-TAC which comprises RNA polymerase, GlnR, and a promoter containing four well-characterized conserved GlnR binding sites. These structures illustrate how four GlnR protomers coordinate to engage promoter DNA in a head-to-tail manner, with four N-terminal receiver domains of GlnR (GlnR-RECs) bridging GlnR_DBDs and the RNAP core enzyme. Structural analysis also unravels that GlnR-TAC is stabilized by complex protein-protein interactions between GlnR and the conserved β flap, σR4, αCTD, and αNTD domains of RNAP, which are further confirmed by our biochemical assays. Taken together, these results reveal a global transcription activation mechanism for the master regulator GlnR and other OmpR/PhoB subfamily proteins and present a unique mode of bacterial transcription regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34816.map.gz emd_34816.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34816-v30.xml emd-34816-v30.xml emd-34816.xml emd-34816.xml | 26.4 KB 26.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34816_fsc.xml emd_34816_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_34816.png emd_34816.png | 58.7 KB | ||
| Filedesc metadata |  emd-34816.cif.gz emd-34816.cif.gz | 8.7 KB | ||
| Others |  emd_34816_half_map_1.map.gz emd_34816_half_map_1.map.gz emd_34816_half_map_2.map.gz emd_34816_half_map_2.map.gz | 23.4 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34816 http://ftp.pdbj.org/pub/emdb/structures/EMD-34816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34816 | HTTPS FTP |
-Related structure data
| Related structure data |  8hihMC  8hmlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34816.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34816.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.19 Å | ||||||||||||||||||||||||||||||||||||
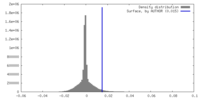
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34816_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
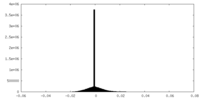
| Density Histograms |
-Half map: #1
| File | emd_34816_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
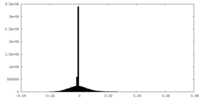
| Density Histograms |
- Sample components
Sample components
+Entire : Mycobacterium tuberculosis transcription initiation complex with ...
+Supramolecule #1: Mycobacterium tuberculosis transcription initiation complex with ...
+Macromolecule #1: DNA-directed RNA polymerase subunit alpha
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase subunit beta'
+Macromolecule #4: DNA-directed RNA polymerase subunit omega
+Macromolecule #5: RNA polymerase sigma factor SigA
+Macromolecule #8: Transcriptional regulatory protein
+Macromolecule #6: Non-template strand DNA of amtB promoter
+Macromolecule #7: Template strand DNA of amtB promoter
+Macromolecule #9: ZINC ION
+Macromolecule #10: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)