+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of human Pannexin isoform 3 | |||||||||
 Map data Map data | Main Map of Pannexin 3 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pannexins / large pore ion-channels / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationgap junction hemi-channel activity / positive regulation of interleukin-1 production / wide pore channel activity / gap junction / monoatomic cation transport / calcium channel activity / osteoblast differentiation / cell-cell signaling / endoplasmic reticulum membrane / structural molecule activity / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.91 Å | |||||||||
 Authors Authors | Hussain N / Penmatsa A | |||||||||
| Funding support |  India, 1 items India, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Cryo-EM structures of pannexin 1 and 3 reveal differences among pannexin isoforms. Authors: Nazia Hussain / Ashish Apotikar / Shabareesh Pidathala / Sourajit Mukherjee / Ananth Prasad Burada / Sujit Kumar Sikdar / Kutti R Vinothkumar / Aravind Penmatsa /   Abstract: Pannexins are single-membrane large-pore channels that release ions and ATP upon activation. Three isoforms of pannexins 1, 2, and 3, perform diverse cellular roles and differ in their pore lining ...Pannexins are single-membrane large-pore channels that release ions and ATP upon activation. Three isoforms of pannexins 1, 2, and 3, perform diverse cellular roles and differ in their pore lining residues. In this study, we report the cryo-EM structure of pannexin 3 at 3.9 Å and analyze its structural differences with pannexin isoforms 1 and 2. The pannexin 3 vestibule has two distinct chambers and a wider pore radius in comparison to pannexins 1 and 2. We further report two cryo-EM structures of pannexin 1, with pore substitutions W74R/R75D that mimic the pore lining residues of pannexin 2 and a germline mutant of pannexin 1, R217H at resolutions of 3.2 Å and 3.9 Å, respectively. Substitution of cationic residues in the vestibule of pannexin 1 results in reduced ATP interaction propensities to the channel. The germline mutant R217H in transmembrane helix 3 (TM3), leads to a partially constricted pore, reduced ATP interaction and weakened voltage sensitivity. The study compares the three pannexin isoform structures, the effects of substitutions of pore and vestibule-lining residues and allosteric effects of a pathological substitution on channel structure and function thereby enhancing our understanding of this vital group of ATP-release channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34265.map.gz emd_34265.map.gz | 168.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34265-v30.xml emd-34265-v30.xml emd-34265.xml emd-34265.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
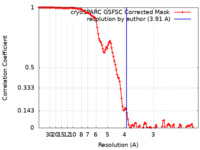
| FSC (resolution estimation) |  emd_34265_fsc.xml emd_34265_fsc.xml | 13.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_34265.png emd_34265.png | 64.9 KB | ||
| Filedesc metadata |  emd-34265.cif.gz emd-34265.cif.gz | 6.7 KB | ||
| Others |  emd_34265_half_map_1.map.gz emd_34265_half_map_1.map.gz emd_34265_half_map_2.map.gz emd_34265_half_map_2.map.gz | 165.2 MB 165.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34265 http://ftp.pdbj.org/pub/emdb/structures/EMD-34265 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34265 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34265 | HTTPS FTP |
-Related structure data
| Related structure data |  8gtrMC  8gtsC  8gttC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34265.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34265.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main Map of Pannexin 3 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.065 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Panx3 halfmapB
| File | emd_34265_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Panx3_halfmapB | ||||||||||||
| Projections & Slices |
| ||||||||||||
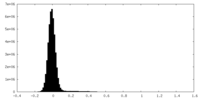
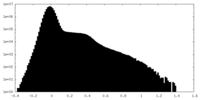
| Density Histograms |
-Half map: Panx3 halfmapA
| File | emd_34265_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Panx3_halfmapA | ||||||||||||
| Projections & Slices |
| ||||||||||||
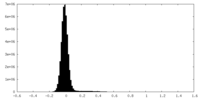
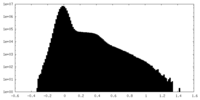
| Density Histograms |
- Sample components
Sample components
-Entire : Pannexin 3
| Entire | Name: Pannexin 3 |
|---|---|
| Components |
|
-Supramolecule #1: Pannexin 3
| Supramolecule | Name: Pannexin 3 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: human isoform 3 of Pannexin. Expressed in ER and Plasma membranes involved in calcium and ATP regulation |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.68 kDa/nm |
-Macromolecule #1: Pannexin-3
| Macromolecule | Name: Pannexin-3 / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.734219 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSLAHTAAEY MLSDALLPDR RGPRLKGLRL ELPLDRIVKF VAVGSPLLLM SLAFAQEFSS GSPISCFSPS NFSIRQAAYV DSSCWDSLL HHKQDGPGQD KMKSLWPHKA LPYSLLALAL LMYLPVLLWQ YAAVPALSSD LLFIISELDK SYNRSIRLVQ H MLKIRQKS ...String: MSLAHTAAEY MLSDALLPDR RGPRLKGLRL ELPLDRIVKF VAVGSPLLLM SLAFAQEFSS GSPISCFSPS NFSIRQAAYV DSSCWDSLL HHKQDGPGQD KMKSLWPHKA LPYSLLALAL LMYLPVLLWQ YAAVPALSSD LLFIISELDK SYNRSIRLVQ H MLKIRQKS SDPYVFWNEL EKARKERYFE FPLLERYLAC KQRSHSLVAT YLLRNSLLLI FTSATYLYLG HFHLDVFFQE EF SCSIKTG LLSDETHVPN LITCRLTSLS IFQIVSLSSV AIYTILVPVI IYNLTRLCRW DKRLLSVYEM LPAFDLLSRK MLG CPINDL NVILLFLRAN ISELISFSWL SVLCVLKDTT TQKHNIDTVV DFMTLLAGLE PSKPKHLTNS ACDEHP UniProtKB: Pannexin-3 |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 7 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: PHOSPHATIDYLETHANOLAMINE
| Macromolecule | Name: PHOSPHATIDYLETHANOLAMINE / type: ligand / ID: 3 / Number of copies: 7 / Formula: PTY |
|---|---|
| Molecular weight | Theoretical: 734.039 Da |
| Chemical component information |  ChemComp-PTY: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Fresh solution containing detergent was prepared for every prep. | |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 288 K / Instrument: FEI VITROBOT MARK IV / Details: blotted for 3.5 seconds. | |||||||||||||||
| Details | Sample is homoheptamer purified to homogeneity. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 100.0 K / Max: 100.0 K |
| Specialist optics | Phase plate: OTHER / Spherical aberration corrector: None / Chromatic aberration corrector: None / Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV / Details: Bioquantum with K2 camera |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 1122 / Average electron dose: 48.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 3.3000000000000003 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 143.6 / Target criteria: Correlation coeficient |
|---|---|
| Output model |  PDB-8gtr: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)