+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Serotonin 4 (5-HT4) receptor-Gi-scFv16 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / serotonin receptor / 5-HT / 5-HT4 receptor / signaling complex / cryo-EM structure / ligand selectivity / G protein selectivity / structure-function relationships / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationlarge intestinal transit / regulation of appetite / maintenance of gastrointestinal epithelium / mucus secretion / serotonin receptor activity / Serotonin receptors / G protein-coupled serotonin receptor activity / neurotransmitter receptor activity / serotonin binding / adenylate cyclase-activating serotonin receptor signaling pathway ...large intestinal transit / regulation of appetite / maintenance of gastrointestinal epithelium / mucus secretion / serotonin receptor activity / Serotonin receptors / G protein-coupled serotonin receptor activity / neurotransmitter receptor activity / serotonin binding / adenylate cyclase-activating serotonin receptor signaling pathway / regulation of postsynapse assembly / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / cellular response to forskolin / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / Regulation of insulin secretion / neuropeptide signaling pathway / response to prostaglandin E / electron transport chain / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / G protein-coupled receptor binding / response to peptide hormone / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / GDP binding / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G alpha (12/13) signalling events / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / sperm principal piece / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / G protein activity / fibroblast proliferation / Ca2+ pathway / midbody / cell cortex / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / chemical synaptic transmission / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / periplasmic space / electron transfer activity / Extra-nuclear estrogen signaling / cell population proliferation / endosome membrane / postsynapse / ciliary basal body / iron ion binding / G protein-coupled receptor signaling pathway Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Huang S / Xu P / Shen DD / Simon IA / Mao C / Tan Y / Zhang H / Harpsoe K / Li H / Zhang Y ...Huang S / Xu P / Shen DD / Simon IA / Mao C / Tan Y / Zhang H / Harpsoe K / Li H / Zhang Y / You C / Yu X / Jiang Y / Gloriam DE / Xu HE | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: GPCRs steer G and G selectivity via TM5-TM6 switches as revealed by structures of serotonin receptors. Authors: Sijie Huang / Peiyu Xu / Dan-Dan Shen / Icaro A Simon / Chunyou Mao / Yangxia Tan / Huibing Zhang / Kasper Harpsøe / Huadong Li / Yumu Zhang / Chongzhao You / Xuekui Yu / Yi Jiang / Yan ...Authors: Sijie Huang / Peiyu Xu / Dan-Dan Shen / Icaro A Simon / Chunyou Mao / Yangxia Tan / Huibing Zhang / Kasper Harpsøe / Huadong Li / Yumu Zhang / Chongzhao You / Xuekui Yu / Yi Jiang / Yan Zhang / David E Gloriam / H Eric Xu /    Abstract: Serotonin (or 5-hydroxytryptamine, 5-HT) is an important neurotransmitter that activates 12 different G protein-coupled receptors (GPCRs) through selective coupling of G, G or G proteins. The ...Serotonin (or 5-hydroxytryptamine, 5-HT) is an important neurotransmitter that activates 12 different G protein-coupled receptors (GPCRs) through selective coupling of G, G or G proteins. The structural basis for G protein subtype selectivity by these GPCRs remains elusive. Here, we report the structures of the serotonin receptors 5-HT, 5-HT, and 5-HT with G, and 5-HT with G. The structures reveal that transmembrane helices TM5 and TM6 alternate lengths as a macro-switch to determine receptor's selectivity for G and G, respectively. We find that the macro-switch by the TM5-TM6 length is shared by class A GPCR-G protein structures. Furthermore, we discover specific residues within TM5 and TM6 that function as micro-switches to form specific interactions with G or G. Together, these results present a common mechanism of G versus G protein coupling selectivity or promiscuity by class A GPCRs and extend the basis of ligand recognition at serotonin receptors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33444.map.gz emd_33444.map.gz | 26.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33444-v30.xml emd-33444-v30.xml emd-33444.xml emd-33444.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33444.png emd_33444.png | 84.2 KB | ||
| Filedesc metadata |  emd-33444.cif.gz emd-33444.cif.gz | 6.6 KB | ||
| Others |  emd_33444_half_map_1.map.gz emd_33444_half_map_1.map.gz emd_33444_half_map_2.map.gz emd_33444_half_map_2.map.gz | 23.2 MB 23.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33444 http://ftp.pdbj.org/pub/emdb/structures/EMD-33444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33444 | HTTPS FTP |
-Related structure data
| Related structure data |  7xtaMC  7xt8C  7xt9C  7xtbC  7xtcC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33444.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33444.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.071 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_33444_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33444_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Serotonin 4 (5-HT4) receptor-Gi-scFv16 complex
| Entire | Name: Serotonin 4 (5-HT4) receptor-Gi-scFv16 complex |
|---|---|
| Components |
|
-Supramolecule #1: Serotonin 4 (5-HT4) receptor-Gi-scFv16 complex
| Supramolecule | Name: Serotonin 4 (5-HT4) receptor-Gi-scFv16 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.414047 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHASM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.915496 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.597682 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MDVQLVESGG GLVQPGGSRK LSCSASGFAF SSFGMHWVRQ APEKGLEWVA YISSGSGTIY YADTVKGRFT ISRDDPKNTL FLQMTSLRS EDTAMYYCVR SIYYYGSSPF DFWGQGTTLT VSSGGGGSGG GGSGGGGSDI VMTQATSSVP VTPGESVSIS C RSSKSLLH ...String: MDVQLVESGG GLVQPGGSRK LSCSASGFAF SSFGMHWVRQ APEKGLEWVA YISSGSGTIY YADTVKGRFT ISRDDPKNTL FLQMTSLRS EDTAMYYCVR SIYYYGSSPF DFWGQGTTLT VSSGGGGSGG GGSGGGGSDI VMTQATSSVP VTPGESVSIS C RSSKSLLH SNGNTYLYWF LQRPGQSPQL LIYRMSNLAS GVPDRFSGSG SGTAFTLTIS RLEAEDVGVY YCMQHLEYPL TF GAGTKLE LK |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: Soluble cytochrome b562,5-hydroxytryptamine receptor 4
| Macromolecule | Name: Soluble cytochrome b562,5-hydroxytryptamine receptor 4 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 59.525551 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: DYKDDDDAKL QTMHHHHHHH HHHADLEDNW ETLNDNLKVI EKADNAAQVK DALTKMRAAA LDAQKATPPK LEDKSPDSPE MKDFRHGFD ILVGQIDDAL KLANEGKVKE AQAAAEQLKT TRNAYIQKYL ASENLYFQGG TDKLDANVSS EEGFGSVEKV V LLTFLSTV ...String: DYKDDDDAKL QTMHHHHHHH HHHADLEDNW ETLNDNLKVI EKADNAAQVK DALTKMRAAA LDAQKATPPK LEDKSPDSPE MKDFRHGFD ILVGQIDDAL KLANEGKVKE AQAAAEQLKT TRNAYIQKYL ASENLYFQGG TDKLDANVSS EEGFGSVEKV V LLTFLSTV ILMAILGNLL VMVAVCWDRQ LRKIKTNYFI VSLAFADLLV SVLVMPFGAI ELVQDIWIYG EVFCLVRTSL DV LLTTASI FHLCCISLDR YYAICCQPLV YRNKMTPLRI ALMLGGCWVI PTFISFLPIM QGWNNIGIID LIEKRKFNQN SNS TYCVFM VNKPYAITCS VVAFYIPFLL MVLAYYRIYV TAKEHAHQIQ MLQRAGASSE SRPQSADQHS THRMRTETKA AKTL CIIMG CFCLCWAPFF VTNIVDPFID YTVPGQVWTA FLWLGYINSG LNPFLYAFLN KSFRRAFLII LCCDDERYRR PSILG QTVP CSTTTINGST HVLRDAVECG GQWESQCHPP ATSPLVAAQP SDT UniProtKB: Soluble cytochrome b562, 5-hydroxytryptamine receptor 4 |
-Macromolecule #6: SEROTONIN
| Macromolecule | Name: SEROTONIN / type: ligand / ID: 6 / Number of copies: 1 / Formula: SRO |
|---|---|
| Molecular weight | Theoretical: 176.215 Da |
| Chemical component information | 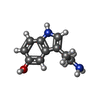 ChemComp-SRO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 611883 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)