[English] 日本語
 Yorodumi
Yorodumi- EMDB-32792: Cryo-EM structure of a yeast pre-40S ribosomal subunit - State Ts... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a yeast pre-40S ribosomal subunit - State Tsr1-1 (with Rps2) | |||||||||
 Map data Map data | local resolution filtered | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome biogenesis / 40S ribosome / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology information18S rRNA (guanine1575-N7)-methyltransferase / rRNA (guanine-N7)-methylation / tRNA methyltransferase complex / rRNA (guanine) methyltransferase activity / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA primary transcript binding / translational readthrough / positive regulation of translational fidelity / : / RMTs methylate histone arginines ...18S rRNA (guanine1575-N7)-methyltransferase / rRNA (guanine-N7)-methylation / tRNA methyltransferase complex / rRNA (guanine) methyltransferase activity / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA primary transcript binding / translational readthrough / positive regulation of translational fidelity / : / RMTs methylate histone arginines / Protein methylation / mTORC1-mediated signalling / Protein hydroxylation / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / Ribosomal scanning and start codon recognition / Major pathway of rRNA processing in the nucleolus and cytosol / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / 90S preribosome / Ub-specific processing proteases / ribosomal subunit export from nucleus / proteasome assembly / regulation of translational fidelity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal small subunit export from nucleus / ribosome assembly / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / small-subunit processome / maintenance of translational fidelity / cytoplasmic stress granule / rRNA processing / : / ribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / cytoplasmic translation / rRNA binding / structural constituent of ribosome / ribosome / translation / mRNA binding / nucleolus / mitochondrion / RNA binding / zinc ion binding / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
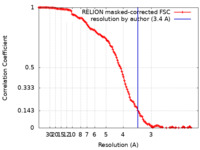
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Cheng J / Lau B / Thoms M / Ameismeier M / Berninghausen O / Hurt E / Beckmann R | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: The nucleoplasmic phase of pre-40S formation prior to nuclear export. Authors: Jingdong Cheng / Benjamin Lau / Matthias Thoms / Michael Ameismeier / Otto Berninghausen / Ed Hurt / Roland Beckmann /   Abstract: Biogenesis of the small ribosomal subunit in eukaryotes starts in the nucleolus with the formation of a 90S precursor and ends in the cytoplasm. Here, we elucidate the enigmatic structural ...Biogenesis of the small ribosomal subunit in eukaryotes starts in the nucleolus with the formation of a 90S precursor and ends in the cytoplasm. Here, we elucidate the enigmatic structural transitions of assembly intermediates from human and yeast cells during the nucleoplasmic maturation phase. After dissociation of all 90S factors, the 40S body adopts a close-to-mature conformation, whereas the 3' major domain, later forming the 40S head, remains entirely immature. A first coordination is facilitated by the assembly factors TSR1 and BUD23-TRMT112, followed by re-positioning of RRP12 that is already recruited early to the 90S for further head rearrangements. Eventually, the uS2 cluster, CK1 (Hrr25 in yeast) and the export factor SLX9 associate with the pre-40S to provide export competence. These exemplary findings reveal the evolutionary conserved mechanism of how yeast and humans assemble the 40S ribosomal subunit, but reveal also a few minor differences. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32792.map.gz emd_32792.map.gz | 105.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32792-v30.xml emd-32792-v30.xml emd-32792.xml emd-32792.xml | 29.4 KB 29.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32792_fsc.xml emd_32792_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_32792.png emd_32792.png | 116.8 KB | ||
| Filedesc metadata |  emd-32792.cif.gz emd-32792.cif.gz | 8.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32792 http://ftp.pdbj.org/pub/emdb/structures/EMD-32792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32792 | HTTPS FTP |
-Related structure data
| Related structure data |  7wtnMC  7wtoC  7wtpC  7wtqC  7wtrC  7wtsC  7wttC  7wtuC  7wtvC  7wtwC  7wtxC  7wtzC  7wu0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32792.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32792.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | local resolution filtered | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Yeast pre-40S ribosomal subunit
+Supramolecule #1: Yeast pre-40S ribosomal subunit
+Macromolecule #1: 18S rRNA
+Macromolecule #2: 40S ribosomal protein S1-A
+Macromolecule #3: 40S ribosomal protein S2
+Macromolecule #4: 40S ribosomal protein S4-A
+Macromolecule #5: 40S ribosomal protein S6-A
+Macromolecule #6: 40S ribosomal protein S7-A
+Macromolecule #7: 40S ribosomal protein S8-A
+Macromolecule #8: 40S ribosomal protein S9-A
+Macromolecule #9: 40S ribosomal protein S11-A
+Macromolecule #10: 40S ribosomal protein S13
+Macromolecule #11: 40S ribosomal protein S14-A
+Macromolecule #12: 40S ribosomal protein S22-A
+Macromolecule #13: 40S ribosomal protein S23-A
+Macromolecule #14: 40S ribosomal protein S24-A
+Macromolecule #15: 40S ribosomal protein S27-A
+Macromolecule #16: 40S ribosomal protein S30-A
+Macromolecule #17: Pre-rRNA-processing protein PNO1
+Macromolecule #18: 18S rRNA (guanine(1575)-N(7))-methyltransferase
+Macromolecule #19: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)