+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of nucleotide-free ABCA3 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC transporter / MEMBRANE PROTEIN / TRANSLOCASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of protein homooligomerization / lamellar body membrane / Defective ABCA3 causes SMDP3 / Defective ABCA3 causes SMDP3 / regulation of phosphatidylcholine metabolic process / alveolar lamellar body membrane / lamellar body / phosphatidylcholine transfer activity / positive regulation of phospholipid transport / alveolar lamellar body ...positive regulation of protein homooligomerization / lamellar body membrane / Defective ABCA3 causes SMDP3 / Defective ABCA3 causes SMDP3 / regulation of phosphatidylcholine metabolic process / alveolar lamellar body membrane / lamellar body / phosphatidylcholine transfer activity / positive regulation of phospholipid transport / alveolar lamellar body / xenobiotic export from cell / organelle assembly / phosphatidylcholine flippase activity / ABC transporters in lipid homeostasis / phosphatidylglycerol metabolic process / phosphatidylcholine metabolic process / regulation of lipid biosynthetic process / positive regulation of phospholipid efflux / lipid transporter activity / phospholipid transport / Surfactant metabolism / phospholipid homeostasis / xenobiotic transmembrane transport / multivesicular body membrane / P-type phospholipid transporter / surfactant homeostasis / ABC-type xenobiotic transporter / ABC-type xenobiotic transporter activity / xenobiotic transport / lipid transport / ATPase-coupled transmembrane transporter activity / positive regulation of cholesterol efflux / lung development / response to glucocorticoid / cytoplasmic vesicle membrane / late endosome / response to xenobiotic stimulus / lysosomal membrane / intracellular membrane-bounded organelle / ATP hydrolysis activity / extracellular space / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Xie T / Zhang ZK / Yue J / Gong X | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Cryo-EM structures of the human surfactant lipid transporter ABCA3. Authors: Tian Xie / Zike Zhang / Jian Yue / Qi Fang / Xin Gong /  Abstract: The adenosine 5'-triphosphate (ATP)-binding cassette (ABC) transporter ABCA3 plays a critical role in pulmonary surfactant biogenesis. Mutations in human ABCA3 have been recognized as the most ...The adenosine 5'-triphosphate (ATP)-binding cassette (ABC) transporter ABCA3 plays a critical role in pulmonary surfactant biogenesis. Mutations in human ABCA3 have been recognized as the most frequent causes of inherited surfactant dysfunction disorders. Despite two decades of research, in vitro biochemical and structural studies of ABCA3 are still lacking. Here, we report the cryo-EM structures of human ABCA3 in two distinct conformations, both at resolution of 3.3 Å. In the absence of ATP, ABCA3 adopts a "lateral-opening" conformation with the lateral surfaces of transmembrane domains (TMDs) exposed to the membrane and features two positively charged cavities within the TMDs as potential substrate binding sites. ATP binding induces pronounced conformational changes, resulting in the collapse of the potential substrate binding cavities. Our results help to rationalize the disease-causing mutations in human ABCA3 and suggest a conserved "lateral access and extrusion" mechanism for both lipid export and import mediated by ABCA transporters. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32233.map.gz emd_32233.map.gz | 49.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32233-v30.xml emd-32233-v30.xml emd-32233.xml emd-32233.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_32233.png emd_32233.png | 31 KB | ||
| Filedesc metadata |  emd-32233.cif.gz emd-32233.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32233 http://ftp.pdbj.org/pub/emdb/structures/EMD-32233 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32233 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32233 | HTTPS FTP |
-Related structure data
| Related structure data |  7w01MC  7w02C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32233.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32233.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : ABCA3
| Entire | Name: ABCA3 |
|---|---|
| Components |
|
-Supramolecule #1: ABCA3
| Supramolecule | Name: ABCA3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Phospholipid-transporting ATPase ABCA3
| Macromolecule | Name: Phospholipid-transporting ATPase ABCA3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: P-type phospholipid transporter |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 196.513656 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RMAVLRQLAL LLWKNYTLQK RKVLVTVLEL FLPLLFSGIL IWLRLKIQSE NVPNATIYPG QSIQELPLF FTFPPPGDTW ELAYIPSHSD AAKTVTETVR RALVINMRVR GFPSEKDFED YIRYDNCSSS VLAAVVFEHP F NHSKEPLP ...String: MADYKDDDDK SGPDEVDASG RMAVLRQLAL LLWKNYTLQK RKVLVTVLEL FLPLLFSGIL IWLRLKIQSE NVPNATIYPG QSIQELPLF FTFPPPGDTW ELAYIPSHSD AAKTVTETVR RALVINMRVR GFPSEKDFED YIRYDNCSSS VLAAVVFEHP F NHSKEPLP LAVKYHLRFS YTRRNYMWTQ TGSFFLKETE GWHTTSLFPL FPNPGPREPT SPDGGEPGYI REGFLAVQHA VD RAIMEYH ADAATRQLFQ RLTVTIKRFP YPPFIADPFL VAIQYQLPLL LLLSFTYTAL TIARAVVQEK ERRLKEYMRM MGL SSWLHW SAWFLLFFLF LLIAASFMTL LFCVKVKPNV AVLSRSDPSL VLAFLLCFAI STISFSFMVS TFFSKANMAA AFGG FLYFF TYIPYFFVAP RYNWMTLSQK LCSCLLSNVA MAMGAQLIGK FEAKGMGIQW RDLLSPVNVD DDFCFGQVLG MLLLD SVLY GLVTWYMEAV FPGQFGVPQP WYFFIMPSYW CGKPRAVAGK EEEDSDPEKA LRNEYFEAEP EDLVAGIKIK HLSKVF RVG NKDRAAVRDL NLNLYEGQIT VLLGHNGAGK TTTLSMLTGL FPPTSGRAYI SGYEISQDMV QIRKSLGLCP QHDILFD NL TVAEHLYFYA QLKGLSRQKC PEEVKQMLHI IGLEDKWNSR SRFLSGGMRR KLSIGIALIA GSKVLILDEP TSGMDAIS R RAIWDLLQRQ KSDRTIVLTT HFMDEADLLG DRIAIMAKGE LQCCGSSLFL KQKYGAGYHM TLVKEPHCNP EDISQLVHH HVPNATLESS AGAELSFILP RESTHRFEGL FAKLEKKQKE LGIASFGASI TTMEEVFLRV GKLVDSSMDI QAIQLPALQY QHERRASDW AVDSNLCGAM DPSDGIGALI EEERTAVKLN TGLALHCQQF WAMFLKKAAY SWREWKMVAA QVLVPLTCVT L ALLAINYS SELFDDPMLR LTLGEYGRTV VPFSVPGTSQ LGQQLSEHLK DALQAEGQEP REVLGDLEEF LIFRASVEGG GF NERCLVA ASFRDVGERT VVNALFNNQA YHSPATALAV VDNLLFKLLC GPHASIVVSN FPQPRSALQA AKDQFNEGRK GFD IALNLL FAMAFLASTF SILAVSERAV QAKHVQFVSG VHVASFWLSA LLWDLISFLI PSLLLLVVFK AFDVRAFTRD GHMA DTLLL LLLYGWAIIP LMYLMNFFFL GAATAYTRLT IFNILSGIAT FLMVTIMRIP AVKLEELSKT LDHVFLVLPN HCLGM AVSS FYENYETRRY CTSSEVAAHY CKKYNIQYQE NFYAWSAPGV GRFVASMAAS GCAYLILLFL IETNLLQRLR GILCAL RRR RTLTELYTRM PVLPEDQDVA DERTRILAPS PDSLLHTPLI IKELSKVYEQ RVPLLAVDRL SLAVQKGECF GLLGFNG AG KTTTFKMLTG EESLTSGDAF VGGHRISSDV GKVRQRIGYC PQFDALLDHM TGREMLVMYA RLRGIPERHI GACVENTL R GLLLEPHANK LVRTYSGGNK RKLSTGIALI GEPAVIFLDE PSTGMDPVAR RLLWDTVARA RESGKAIIIT SHSMEECEA LCTRLAIMVQ GQFKCLGSPQ HLKSKFGSGY SLRAKVQSEG QQEALEEFKA FVDLTFPGSV LEDEHQGMVH YHLPGRDLSW AKVFGILEK AKEKYGVDDY SVSQISLEQV FLSFAHLQPP TAEEGRLEGS DEVDAVEGSH HHHHHHHHH UniProtKB: Phospholipid-transporting ATPase ABCA3 |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 3 / Number of copies: 2 / Formula: PX4 |
|---|---|
| Molecular weight | Theoretical: 678.94 Da |
| Chemical component information | 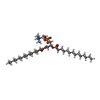 ChemComp-PX4: |
-Macromolecule #4: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 4 / Number of copies: 3 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




















