[English] 日本語
 Yorodumi
Yorodumi- EMDB-32227: Cryo-EM structure of amyloid fibril formed by full-length human SOD1 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of amyloid fibril formed by full-length human SOD1 | ||||||||||||
 Map data Map data | The cryo-EM map of SOD1 fibril. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Amyloid fibril / PROTEIN FIBRIL | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationaction potential initiation / response to antipsychotic drug / neurofilament cytoskeleton organization / response to carbon monoxide / relaxation of vascular associated smooth muscle / dense core granule / regulation of organ growth / response to superoxide / positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / peripheral nervous system myelin maintenance ...action potential initiation / response to antipsychotic drug / neurofilament cytoskeleton organization / response to carbon monoxide / relaxation of vascular associated smooth muscle / dense core granule / regulation of organ growth / response to superoxide / positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / peripheral nervous system myelin maintenance / anterograde axonal transport / protein phosphatase 2B binding / regulation of T cell differentiation in thymus / Oxidoreductases; Acting on a sulfur group of donors / regulation of GTPase activity / retina homeostasis / auditory receptor cell stereocilium organization / cellular response to potassium ion / hydrogen peroxide biosynthetic process / retrograde axonal transport / myeloid cell homeostasis / superoxide anion generation / superoxide metabolic process / response to copper ion / superoxide dismutase / muscle cell cellular homeostasis / Detoxification of Reactive Oxygen Species / superoxide dismutase activity / heart contraction / cellular response to ATP / cellular response to cadmium ion / negative regulation of reproductive process / negative regulation of developmental process / transmission of nerve impulse / regulation of multicellular organism growth / ectopic germ cell programmed cell death / response to axon injury / ovarian follicle development / neuronal action potential / positive regulation of superoxide anion generation / axon cytoplasm / removal of superoxide radicals / embryo implantation / reactive oxygen species metabolic process / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / positive regulation of phagocytosis / dendrite cytoplasm / placenta development / thymus development / positive regulation of cytokine production / regulation of mitochondrial membrane potential / determination of adult lifespan / response to amphetamine / glutathione metabolic process / response to hydrogen peroxide / locomotory behavior / sensory perception of sound / response to nutrient levels / mitochondrial intermembrane space / negative regulation of inflammatory response / regulation of blood pressure / small GTPase binding / Platelet degranulation / peroxisome / response to heat / protein-folding chaperone binding / cytoplasmic vesicle / spermatogenesis / gene expression / negative regulation of neuron apoptotic process / intracellular iron ion homeostasis / response to ethanol / positive regulation of MAPK cascade / lysosome / positive regulation of apoptotic process / response to xenobiotic stimulus / mitochondrial matrix / copper ion binding / neuronal cell body / apoptotic process / protein homodimerization activity / protein-containing complex / mitochondrion / : / extracellular exosome / extracellular region / zinc ion binding / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
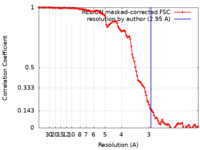
| Method | helical reconstruction / cryo EM / Resolution: 2.95 Å | ||||||||||||
 Authors Authors | Wang LQ / Ma YY / Yuan HY / Zhao K / Zhang MY / Wang Q / Huang X / Xu WC / Chen J / Li D ...Wang LQ / Ma YY / Yuan HY / Zhao K / Zhang MY / Wang Q / Huang X / Xu WC / Chen J / Li D / Zhang DL / Zou LY / Yin P / Liu C / Liang Y | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Cryo-EM structure of an amyloid fibril formed by full-length human SOD1 reveals its conformational conversion. Authors: Li-Qiang Wang / Yeyang Ma / Han-Ye Yuan / Kun Zhao / Mu-Ya Zhang / Qiang Wang / Xi Huang / Wen-Chang Xu / Bin Dai / Jie Chen / Dan Li / Delin Zhang / Zhengzhi Wang / Liangyu Zou / Ping Yin / ...Authors: Li-Qiang Wang / Yeyang Ma / Han-Ye Yuan / Kun Zhao / Mu-Ya Zhang / Qiang Wang / Xi Huang / Wen-Chang Xu / Bin Dai / Jie Chen / Dan Li / Delin Zhang / Zhengzhi Wang / Liangyu Zou / Ping Yin / Cong Liu / Yi Liang /  Abstract: Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease. Misfolded Cu, Zn-superoxide dismutase (SOD1) has been linked to both familial and sporadic ALS. SOD1 fibrils formed in vitro share ...Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease. Misfolded Cu, Zn-superoxide dismutase (SOD1) has been linked to both familial and sporadic ALS. SOD1 fibrils formed in vitro share toxic properties with ALS inclusions. Here we produced cytotoxic amyloid fibrils from full-length apo human SOD1 under reducing conditions and determined the atomic structure using cryo-EM. The SOD1 fibril consists of a single protofilament with a left-handed helix. The fibril core exhibits a serpentine fold comprising N-terminal segment (residues 3-55) and C-terminal segment (residues 86-153) with an intrinsic disordered segment. The two segments are zipped up by three salt bridge pairs. By comparison with the structure of apo SOD1 dimer, we propose that eight β-strands (to form a β-barrel) and one α-helix in the subunit of apo SOD1 convert into thirteen β-strands stabilized by five hydrophobic cavities in the SOD1 fibril. Our data provide insights into how SOD1 converts between structurally and functionally distinct states. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32227.map.gz emd_32227.map.gz | 9.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32227-v30.xml emd-32227-v30.xml emd-32227.xml emd-32227.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32227_fsc.xml emd_32227_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_32227.png emd_32227.png | 100.7 KB | ||
| Filedesc metadata |  emd-32227.cif.gz emd-32227.cif.gz | 5.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32227 http://ftp.pdbj.org/pub/emdb/structures/EMD-32227 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32227 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32227 | HTTPS FTP |
-Related structure data
| Related structure data |  7vzfMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32227.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32227.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The cryo-EM map of SOD1 fibril. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Human SOD1 amyloid fibril
| Entire | Name: Human SOD1 amyloid fibril |
|---|---|
| Components |
|
-Supramolecule #1: Human SOD1 amyloid fibril
| Supramolecule | Name: Human SOD1 amyloid fibril / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Superoxide dismutase [Cu-Zn]
| Macromolecule | Name: Superoxide dismutase [Cu-Zn] / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: superoxide dismutase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.958757 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MATKAVCVLK GDGPVQGIIN FEQKESNGPV KVWGSIKGLT EGLHGFHVHE FGDNTAGCTS AGPHFNPLSR KHGGPKDEER HVGDLGNVT ADKDGVADVS IEDSVISLSG DHCIIGRTLV VHEKADDLGK GGNEESTKTG NAGSRLACGV IGIAQ UniProtKB: Superoxide dismutase [Cu-Zn] |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)