+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-32120 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of mouse NLRP3 (full-length) dodecamer | |||||||||||||||
 Map data Map data | cryo-EM map of mouse NLRP3 dodecamer | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | NLR / NOD-like receptor / NLRP3 / Inflammasome / IMMUNE SYSTEM | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of T-helper 17 cell differentiation / The NLRP3 inflammasome / detection of biotic stimulus / Metalloprotease DUBs / molecular sensor activity / phosphatidylinositol phosphate binding / positive regulation of T-helper 2 cell differentiation / canonical inflammasome complex / positive regulation of T-helper 2 cell cytokine production / acute inflammatory response ...positive regulation of T-helper 17 cell differentiation / The NLRP3 inflammasome / detection of biotic stimulus / Metalloprotease DUBs / molecular sensor activity / phosphatidylinositol phosphate binding / positive regulation of T-helper 2 cell differentiation / canonical inflammasome complex / positive regulation of T-helper 2 cell cytokine production / acute inflammatory response / positive regulation of interleukin-18 production / interphase microtubule organizing center / positive regulation of type 2 immune response / positive regulation of cytokine production involved in immune response / NLRP3 inflammasome complex / NLRP3 inflammasome complex assembly / cellular response to peptidoglycan / positive regulation of interleukin-5 production / positive regulation of interleukin-13 production / interleukin-18-mediated signaling pathway / cysteine-type endopeptidase activator activity / phosphatidylinositol-4-phosphate binding / osmosensory signaling pathway / negative regulation of non-canonical NF-kappaB signal transduction / interleukin-1-mediated signaling pathway / negative regulation of interleukin-1 beta production / pattern recognition receptor signaling pathway / positive regulation of interleukin-4 production / negative regulation of acute inflammatory response / positive regulation of NLRP3 inflammasome complex assembly / microtubule organizing center / protein secretion / pyroptotic inflammatory response / signaling adaptor activity / positive regulation of interleukin-1 beta production / molecular condensate scaffold activity / protein maturation / ADP binding / cellular response to virus / protein homooligomerization / positive regulation of non-canonical NF-kappaB signal transduction / negative regulation of inflammatory response / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / positive regulation of inflammatory response / cellular response to lipopolysaccharide / regulation of inflammatory response / DNA-binding transcription factor binding / sequence-specific DNA binding / defense response to virus / molecular adaptor activity / response to ethanol / protein-macromolecule adaptor activity / defense response to Gram-positive bacterium / inflammatory response / Golgi membrane / innate immune response / endoplasmic reticulum / ATP hydrolysis activity / positive regulation of transcription by RNA polymerase II / mitochondrion / extracellular region / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.55 Å | |||||||||||||||
 Authors Authors | Ohto U / Shimizu T | |||||||||||||||
| Funding support |  Japan, 4 items Japan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Structural basis for the oligomerization-mediated regulation of NLRP3 inflammasome activation. Authors: Umeharu Ohto / Yukie Kamitsukasa / Hanako Ishida / Zhikuan Zhang / Karin Murakami / Chie Hirama / Sakiko Maekawa / Toshiyuki Shimizu /  Abstract: SignificanceThe nucleotide-binding oligomerization domain (NOD)-like receptor pyrin domain containing 3 (NLRP3) is a pattern recognition receptor that forms an inflammasome. The cryo-electron ...SignificanceThe nucleotide-binding oligomerization domain (NOD)-like receptor pyrin domain containing 3 (NLRP3) is a pattern recognition receptor that forms an inflammasome. The cryo-electron microscopy structure of the dodecameric form of full-length NLRP3 bound to the clinically relevant NLRP3-specific inhibitor MCC950 has established the structural basis for the oligomerization-mediated regulation of NLRP3 inflammasome activation and the mechanism of action of the NLRP3 specific inhibitor. The inactive NLRP3 oligomer represents the NLRP3 resting state, capable of binding to membranes and is likely disrupted for its activation. Visualization of the inhibitor binding mode will enable optimization of the activity of NLRP3 inflammasome inhibitor drugs. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32120.map.gz emd_32120.map.gz | 39.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32120-v30.xml emd-32120-v30.xml emd-32120.xml emd-32120.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_32120.png emd_32120.png | 144.7 KB | ||
| Filedesc metadata |  emd-32120.cif.gz emd-32120.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32120 http://ftp.pdbj.org/pub/emdb/structures/EMD-32120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32120 | HTTPS FTP |
-Related structure data
| Related structure data |  7vtqMC  7vtpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32120.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32120.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM map of mouse NLRP3 dodecamer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.245 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mouse NLRP3 (full-length) dodecamer
| Entire | Name: Mouse NLRP3 (full-length) dodecamer |
|---|---|
| Components |
|
-Supramolecule #1: Mouse NLRP3 (full-length) dodecamer
| Supramolecule | Name: Mouse NLRP3 (full-length) dodecamer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: NACHT, LRR and PYD domains-containing protein 3
| Macromolecule | Name: NACHT, LRR and PYD domains-containing protein 3 / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 121.415977 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HHHHHHDYKD DDDKLEVLFQ GPEFMTSVRC KLAQYLEDLE DVDLKKFKMH LEDYPPEKGC IPVPRGQMEK ADHLDLATLM IDFNGEEKA WAMAVWIFAA INRRDLWEKA KKDQPEWNDT CTSHSSMVCQ EDSLEEEWMG LLGYLSRISI CKKKKDYCKM Y RRHVRSRF ...String: HHHHHHDYKD DDDKLEVLFQ GPEFMTSVRC KLAQYLEDLE DVDLKKFKMH LEDYPPEKGC IPVPRGQMEK ADHLDLATLM IDFNGEEKA WAMAVWIFAA INRRDLWEKA KKDQPEWNDT CTSHSSMVCQ EDSLEEEWMG LLGYLSRISI CKKKKDYCKM Y RRHVRSRF YSIKDRNARL GESVDLNSRY TQLQLVKEHP SKQEREHELL TIGRTKMRDS PMSSLKLELL FEPEDGHSEP VH TVVFQGA AGIGKTILAR KIMLDWALGK LFKDKFDYLF FIHCREVSLR TPRSLADLIV SCWPDPNPPV CKILRKPSRI LFL MDGFDE LQGAFDEHIG EVCTDWQKAV RGDILLSSLI RKKLLPKASL LITTRPVALE KLQHLLDHPR HVEILGFSEA KRKE YFFKY FSNELQAREA FRLIQENEVL FTMCFIPLVC WIVCTGLKQQ METGKSLAQT SKTTTAVYVF FLSSLLQSRG GIEEH LFSD YLQGLCSLAA DGIWNQKILF EECDLRKHGL QKTDVSAFLR MNVFQKEVDC ERFYSFSHMT FQEFFAAMYY LLEEEA EGE TVRKGPGGCS DLLNRDVKVL LENYGKFEKG YLIFVVRFLF GLVNQERTSY LEKKLSCKIS QQVRLELLKW IEVKAKA KK LQWQPSQLEL FYCLYEMQEE DFVQSAMDHF PKIEINLSTR MDHVVSSFCI KNCHRVKTLS LGFFHNSPKE EEEERRGG R PLDQVQCVFP DTHVACSSRL VNCCLTSSFC RGLFSSLSTN RSLTELDLSD NTLGDPGMRV LCEALQHPGC NIQRLWLGR CGLSHQCCFD ISSVLSSSQK LVELDLSDNA LGDFGIRLLC VGLKHLLCNL QKLWLVSCCL TSACCQDLAL VLSSNHSLTR LYIGENALG DSGVQVLCEK MKDPQCNLQK LGLVNSGLTS ICCSALTSVL KTNQNFTHLY LRSNALGDTG LRLLCEGLLH P DCKLQMLE LDNCSLTSHS CWNLSTILTH NHSLRKLNLG NNDLGDLCVV TLCEVLKQQG CLLQSLQLGE MYLNRETKRA LE ALQEEKP ELTIVFEISW UniProtKB: NACHT, LRR and PYD domains-containing protein 3 |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 12 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: 1-[4-(2-oxidanylpropan-2-yl)furan-2-yl]sulfonyl-3-(1,2,3,5-tetrah...
| Macromolecule | Name: 1-[4-(2-oxidanylpropan-2-yl)furan-2-yl]sulfonyl-3-(1,2,3,5-tetrahydro-s-indacen-4-yl)urea type: ligand / ID: 3 / Number of copies: 12 / Formula: 7YN |
|---|---|
| Molecular weight | Theoretical: 402.464 Da |
| Chemical component information | 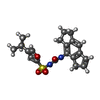 ChemComp-7YN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 25 mM HEPES-NaOH (pH 7.5), 0.2 M NaCl, 1 mM MgCl2, 0.5 mM TCEP, 1.0 mM ADP, 0.05 mM MCC950 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 68.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D6 (2x6 fold dihedral) / Resolution.type: BY AUTHOR / Resolution: 3.55 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 73930 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















