[English] 日本語
 Yorodumi
Yorodumi- EMDB-12582: CryoEM structure of Nipah virus nucleocapsid semi-spiral clam-sha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12582 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Nipah virus nucleocapsid semi-spiral clam-shaped assembly | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative stranded viral RNA transcription / negative stranded viral RNA replication / helical viral capsid / viral nucleocapsid / molecular adaptor activity / host cell cytoplasm / ribonucleoprotein complex / structural molecule activity / RNA binding Similarity search - Function | |||||||||
| Biological species |  Nipah henipavirus Nipah henipavirus | |||||||||
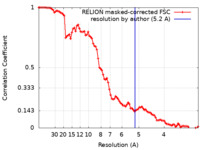
| Method | single particle reconstruction / cryo EM / Resolution: 5.2 Å | |||||||||
 Authors Authors | Ker DS / Jenkins HT / Greive SJ / Antson AA | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2021 Journal: PLoS Pathog / Year: 2021Title: CryoEM structure of the Nipah virus nucleocapsid assembly. Authors: De-Sheng Ker / Huw T Jenkins / Sandra J Greive / Alfred A Antson /  Abstract: Nipah and its close relative Hendra are highly pathogenic zoonotic viruses, storing their ssRNA genome in a helical nucleocapsid assembly formed by the N protein, a major viral immunogen. Here, we ...Nipah and its close relative Hendra are highly pathogenic zoonotic viruses, storing their ssRNA genome in a helical nucleocapsid assembly formed by the N protein, a major viral immunogen. Here, we report the first cryoEM structure for a Henipavirus RNA-bound nucleocapsid assembly, at 3.5 Å resolution. The helical assembly is stabilised by previously undefined N- and C-terminal segments, contributing to subunit-subunit interactions. RNA is wrapped around the nucleocapsid protein assembly with a periodicity of six nucleotides per protomer, in the "3-bases-in, 3-bases-out" conformation, with protein plasticity enabling non-sequence specific interactions. The structure reveals commonalities in RNA binding pockets and in the conformation of bound RNA, not only with members of the Paramyxoviridae family, but also with the evolutionarily distant Filoviridae Ebola virus. Significant structural differences with other Paramyxoviridae members are also observed, particularly in the position and length of the exposed α-helix, residues 123-139, which may serve as a valuable epitope for surveillance and diagnostics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12582.map.gz emd_12582.map.gz | 28.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12582-v30.xml emd-12582-v30.xml emd-12582.xml emd-12582.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12582_fsc.xml emd_12582_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_12582.png emd_12582.png | 68.9 KB | ||
| Masks |  emd_12582_msk_1.map emd_12582_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Others |  emd_12582_half_map_1.map.gz emd_12582_half_map_1.map.gz emd_12582_half_map_2.map.gz emd_12582_half_map_2.map.gz | 27.4 MB 27.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12582 http://ftp.pdbj.org/pub/emdb/structures/EMD-12582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12582 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12582.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12582.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.572 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12582_msk_1.map emd_12582_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_12582_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_12582_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Nipah virus nucleocapsid Protein-RNA complex
| Entire | Name: Nipah virus nucleocapsid Protein-RNA complex |
|---|---|
| Components |
|
-Supramolecule #1: Nipah virus nucleocapsid Protein-RNA complex
| Supramolecule | Name: Nipah virus nucleocapsid Protein-RNA complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Nipah henipavirus Nipah henipavirus |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1 MDa |
-Macromolecule #1: Nucleocapsid protein
| Macromolecule | Name: Nucleocapsid protein / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Nipah henipavirus Nipah henipavirus |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLEVLFQG PAMSDIFEEA ASFRSYQSKL GRDGRASAAT ATLTTKIRI FVPATNSPEL RWELTLFALD VIRSPSAAES MKVGAAFTLI S MYSERPGA LIRSLLNDPD IEAVIIDVGS MVNGIPVMER RGDKAQEEME GL MRILKTA RDSSKGKTPF ...String: MGSSHHHHHH SSGLEVLFQG PAMSDIFEEA ASFRSYQSKL GRDGRASAAT ATLTTKIRI FVPATNSPEL RWELTLFALD VIRSPSAAES MKVGAAFTLI S MYSERPGA LIRSLLNDPD IEAVIIDVGS MVNGIPVMER RGDKAQEEME GL MRILKTA RDSSKGKTPF VDSRAYGLRI TDMSTLVSAV ITIEAQIWIL IAK AVTAPD TAEESETRRW AKYVQQKRVN PFFALTQQWL TEMRNLLSQS LSVR KFMVE ILIEVKKGGS AKGRAVEIIS DIGNYVEETG MAGFFATIRF GLETR YPAL ALNEFQSDLN TIKSLMLLYR EIGPRAPYMV LLEESIQTKF APGGYP LLW SFAMGVATTI DRSMGALNIN RGYLEPMYFR LGQKSARHHA GGIDQNM AN RLGLSSDQVA ELAAAVQETS AGRQESNVQA REAKFAAGGV LIGGSDQD I DEGEEPIEQS GRQSVTFKRE MSISSLANSV PSSSVSTSGG TRLTNSLLN LRSRLAAKAA KEAASSNATD DPAISNRTQG ESEKKNNQDL KPAQNDLDFV RADV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 41.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)