[English] 日本語
 Yorodumi
Yorodumi- EMDB-3033: Structure of PhnGHIJK complex by negative stain electron microscopy -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3033 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of PhnGHIJK complex by negative stain electron microscopy | |||||||||
 Map data Map data | 3D reconstruction of C-P lyase core complex including K subunit | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phosphorus metabolism / carbon-phosphorous lyase / phosphonate / PhnG / PhnH / PhnI / PhnJ / PhnK | |||||||||
| Biological species |  | |||||||||
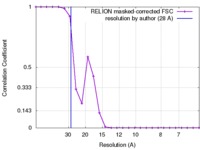
| Method | single particle reconstruction / negative staining / Resolution: 16.0 Å | |||||||||
 Authors Authors | Seweryn P / Bich Van L / Kjeldgaard M / Russo CJ / Passmore LA / Hove-Jensen B / Jochimsen B / Brodersen DE | |||||||||
 Citation Citation |  Journal: Nature / Year: 2015 Journal: Nature / Year: 2015Title: Structural insights into the bacterial carbon-phosphorus lyase machinery. Authors: Paulina Seweryn / Lan Bich Van / Morten Kjeldgaard / Christopher J Russo / Lori A Passmore / Bjarne Hove-Jensen / Bjarne Jochimsen / Ditlev E Brodersen /   Abstract: Phosphorus is required for all life and microorganisms can extract it from their environment through several metabolic pathways. When phosphate is in limited supply, some bacteria are able to use ...Phosphorus is required for all life and microorganisms can extract it from their environment through several metabolic pathways. When phosphate is in limited supply, some bacteria are able to use phosphonate compounds, which require specialized enzymatic machinery to break the stable carbon-phosphorus (C-P) bond. Despite its importance, the details of how this machinery catabolizes phosphonates remain unknown. Here we determine the crystal structure of the 240-kilodalton Escherichia coli C-P lyase core complex (PhnG-PhnH-PhnI-PhnJ; PhnGHIJ), and show that it is a two-fold symmetric hetero-octamer comprising an intertwined network of subunits with unexpected self-homologies. It contains two potential active sites that probably couple phosphonate compounds to ATP and subsequently hydrolyse the C-P bond. We map the binding site of PhnK on the complex using electron microscopy, and show that it binds to a conserved insertion domain of PhnJ. Our results provide a structural basis for understanding microbial phosphonate breakdown. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3033.map.gz emd_3033.map.gz | 253.2 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3033-v30.xml emd-3033-v30.xml emd-3033.xml emd-3033.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3033_fsc.xml emd_3033_fsc.xml | 2 KB | Display |  FSC data file FSC data file |
| Images |  EMD-3033.png EMD-3033.png | 55 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3033 http://ftp.pdbj.org/pub/emdb/structures/EMD-3033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3033 | HTTPS FTP |
-Validation report
| Summary document |  emd_3033_validation.pdf.gz emd_3033_validation.pdf.gz | 226.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3033_full_validation.pdf.gz emd_3033_full_validation.pdf.gz | 225.9 KB | Display | |
| Data in XML |  emd_3033_validation.xml.gz emd_3033_validation.xml.gz | 6.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3033 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3033 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3033 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3033 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3033.map.gz / Format: CCP4 / Size: 670.9 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3033.map.gz / Format: CCP4 / Size: 670.9 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D reconstruction of C-P lyase core complex including K subunit | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Escherichia coli PhnGHIJK complex
| Entire | Name: Escherichia coli PhnGHIJK complex |
|---|---|
| Components |
|
-Supramolecule #1000: Escherichia coli PhnGHIJK complex
| Supramolecule | Name: Escherichia coli PhnGHIJK complex / type: sample / ID: 1000 / Number unique components: 5 |
|---|---|
| Molecular weight | Theoretical: 268 KDa |
-Macromolecule #1: PhnG
| Macromolecule | Name: PhnG / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Oligomeric state: heterodimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: PhnH
| Macromolecule | Name: PhnH / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Oligomeric state: heterodimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #3: PhnI
| Macromolecule | Name: PhnI / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Oligomeric state: heterodimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #4: PhnJ
| Macromolecule | Name: PhnJ / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Oligomeric state: heterodimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 32 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #5: PhnK
| Macromolecule | Name: PhnK / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 28 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Staining | Type: NEGATIVE Details: 3% ammonium molybdate pH 8.0 followed by 2% uranyl acetate |
|---|---|
| Grid | Details: Quantifoil R2/2 holey carbon on copper mesh grids, covered with an additional thin film of amorphous carbon |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Date | Jul 26, 2013 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 105 / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.974 µm / Nominal defocus min: 0.832 µm / Nominal magnification: 44000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)