+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3024 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the type IV pilus from Thermus thermophilus | |||||||||
 Map data Map data | Subtomogram average of the Thermus thermophilus type IV pilus. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Electron cryo-tomography / subtomogram averaging / type IV pilus / DNA transporter / bacterial secretion / T. thermophilus | |||||||||
| Biological species |   Thermus thermophilus HB27 (bacteria) Thermus thermophilus HB27 (bacteria) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 32.0 Å | |||||||||
 Authors Authors | Gold VAM / Salzer R / Averhoff B / Kuehlbrandt W | |||||||||
 Citation Citation |  Journal: Elife / Year: 2015 Journal: Elife / Year: 2015Title: Structure of a type IV pilus machinery in the open and closed state. Authors: Vicki A M Gold / Ralf Salzer / Beate Averhoff / Werner Kühlbrandt /  Abstract: Proteins of the secretin family form large macromolecular complexes, which assemble in the outer membrane of Gram-negative bacteria. Secretins are major components of type II and III secretion ...Proteins of the secretin family form large macromolecular complexes, which assemble in the outer membrane of Gram-negative bacteria. Secretins are major components of type II and III secretion systems and are linked to extrusion of type IV pili (T4P) and to DNA uptake. By electron cryo-tomography of whole Thermus thermophilus cells, we determined the in situ structure of a T4P molecular machine in the open and the closed state. Comparison reveals a major conformational change whereby the N-terminal domains of the central secretin PilQ shift by ~30 Å, and two periplasmic gates open to make way for pilus extrusion. Furthermore, we determine the structure of the assembled pilus. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3024.map.gz emd_3024.map.gz | 141.8 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3024-v30.xml emd-3024-v30.xml emd-3024.xml emd-3024.xml | 8.9 KB 8.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3024.tif emd_3024.tif | 754.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3024 http://ftp.pdbj.org/pub/emdb/structures/EMD-3024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3024 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3024.map.gz / Format: CCP4 / Size: 188.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3024.map.gz / Format: CCP4 / Size: 188.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of the Thermus thermophilus type IV pilus. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.209 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
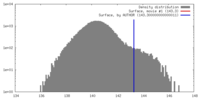
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Type IV pilus
| Entire | Name: Type IV pilus |
|---|---|
| Components |
|
-Supramolecule #1000: Type IV pilus
| Supramolecule | Name: Type IV pilus / type: sample / ID: 1000 Oligomeric state: Thousands of copies of PilA4 form the type IV pilus Number unique components: 1 |
|---|
-Macromolecule #1: Type IV pilus
| Macromolecule | Name: Type IV pilus / type: protein_or_peptide / ID: 1 / Name.synonym: T4P / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB27 (bacteria) Thermus thermophilus HB27 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 20 mM Tris, 100 mM EDTA |
|---|---|
| Grid | Details: 300 mesh copper grid with quantifoil support film (R2/2), glow discharged |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 20 % / Instrument: HOMEMADE PLUNGER Method: Cell solutions were mixed 1:1 (v/v) with 10 nm ProteinA-gold particles. 3 microlitres of sample were applied to grids and blotted on one side for ~5s before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected on the K2 camera at magnification used for imaging |
| Specialist optics | Energy filter - Name: GIF Quantum / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Dec 17, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Average electron dose: 140 e/Å2 Details: Each image in every tilt series is a drift corrected sum of 3-5 frames |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 11640 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 5.0 µm / Nominal magnification: 33000 |
| Sample stage | Specimen holder: Nitrogen cooled / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Subvolumes of 2 nm length were selected along the pili |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 32.0 Å / Resolution method: OTHER / Software - Name: IMOD, (ETOMO, &, PEET), Spider / Number subtomograms used: 740 |
| CTF correction | Details: CTF correction of each tilt image using IMOD |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)